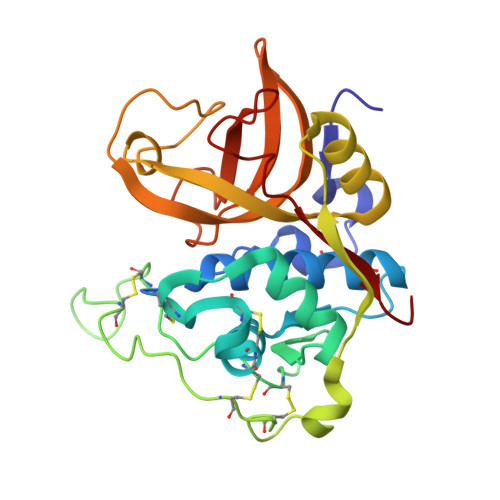
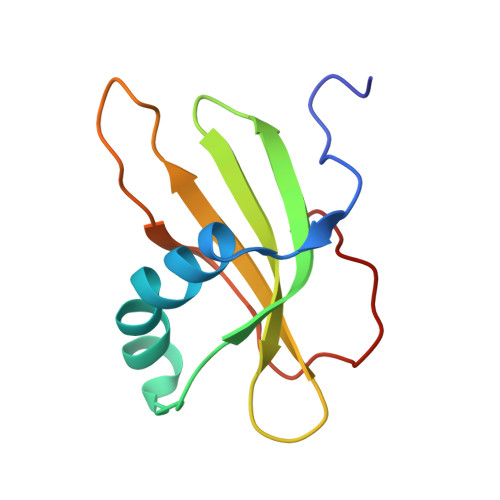
Stefin A displaces the occluding loop of cathepsin B only by as much as required to bind to the active site cleft
Renko, M., Pozgan, U., Majera, D., Turk, D.(2010) FEBS J 277: 4338-4345
- PubMed: 20860624 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07824.x
- Primary Citation Related Structures:
3K9M - PubMed Abstract:
Cathepsin B (EC 3.4.22.1) is one of the most versatile human cysteine cathepsins. It is important for intracellular protein degradation under normal conditions and is involved in a number of pathological processes. The occluding loop makes cathepsin B unique among cysteine cathepsins. This ∼ 20 residue long insertion imbedded into the papain-like protease scaffold restricts access to the active site cleft and endows cathepsin B with its carboxydipeptidase activity. Nevertheless, the enzyme also exhibits endopeptidase activity and is inhibited by stefins and cystatins. To clarify the structural properties of the occluding loop upon the binding of stefins, we determined the crystal structure of the complex between wild-type human stefin A and wild-type human cathepsin B at 2.6 Å resolution. The papain-like part of cathepsin B structure remains unmodified, whereas the occluding loop residues are displaced. The part enclosed by the disulfide bridge containing histidines 110 and 111 (i.e. the 'lasso' part) is rotated by ∼ 45° away from its original position. A comparison of the structure of the unliganded cathepsin B with the structure of the proenzyme, its complexes with chagasin and stefin A shows that the magnitude of the shift of the occluding loop is related to the size of the binding region. It is smallest in the procathepsin structures and increases in the series of complexes with stefin A and chagasin, although it has no impact on the binding constant. Hence, cathepsin B can dock inhibitors and certain substrates regardless of the size of the binding region.
- Department of Biochemistry and Molecular and Structural Biology, Jozef Stefan Institute, Ljubljana, Slovenia.
Organizational Affiliation: