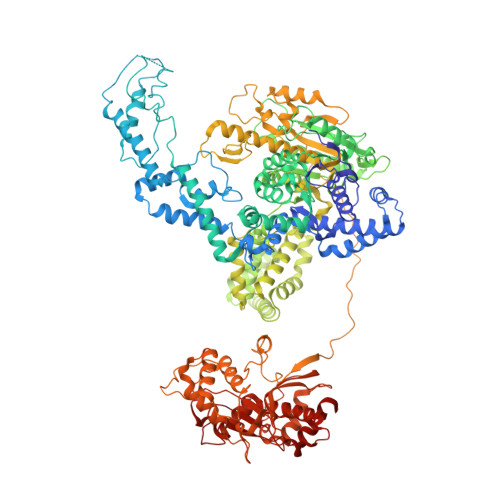
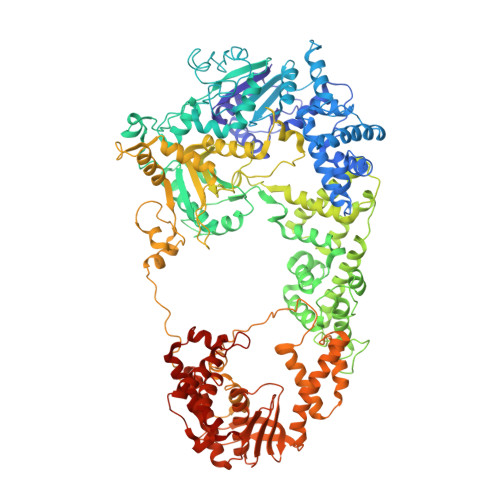
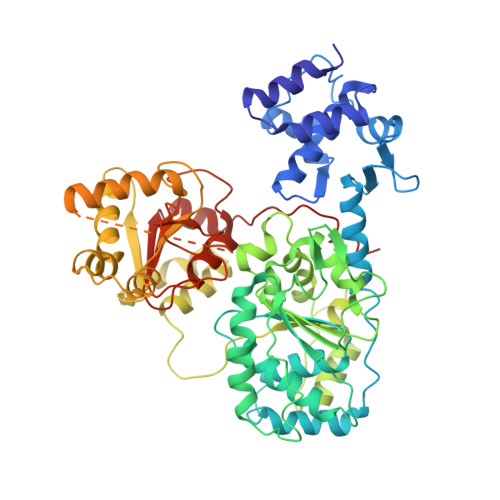
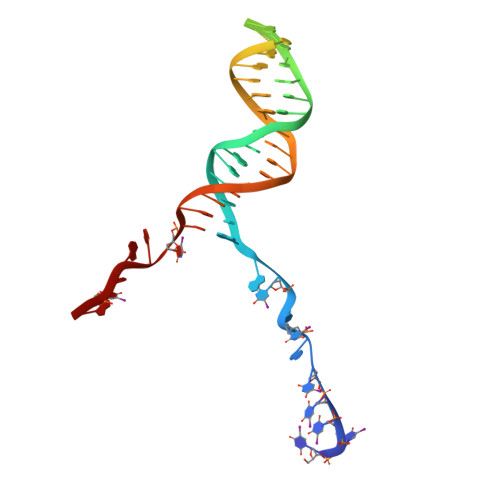
DNA binding to RecD: role of the 1B domain in SF1B helicase activity.
Saikrishnan, K., Griffiths, S.P., Cook, N., Court, R., Wigley, D.B.(2008) EMBO J 27: 2222-2229
- PubMed: 18668125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2008.144
- Primary Citation Related Structures:
3E1S, 3K70 - PubMed Abstract:
The molecular mechanism of superfamily 1Balpha helicases remains unclear. We present here the crystal structure of the RecD2 helicase from Deinococcus radiodurans at 2.2-A resolution. The structure reveals the folds of the 1B and 2B domains of RecD that were poorly ordered in the structure of the Escherichia coli RecBCD enzyme complex reported previously. The 2B domain adopts an SH3 fold which, although common in eukaryotes, is extremely rare in bacterial systems. In addition, the D. radiodurans RecD2 structure has aided us in deciphering lower resolution (3.6 A) electron density maps for the E. coli RecBCD enzyme in complex with a long DNA substrate that interacts with the RecD subunit. Taken together, these structures indicated an important role for the 1B domain of RecD, a beta-hairpin that extends from the surface of the 1A domain and interacts with the DNA substrate. On the basis of these structural data, we designed a mutant RecD2 helicase that lacks this pin. The 'pin-less' mutant protein is a fully active ssDNA-dependent ATPase but totally lacks helicase activity.
- Cancer Research UK Clare Hall Laboratories, The London Research Institute, Herts, UK.
Organizational Affiliation: