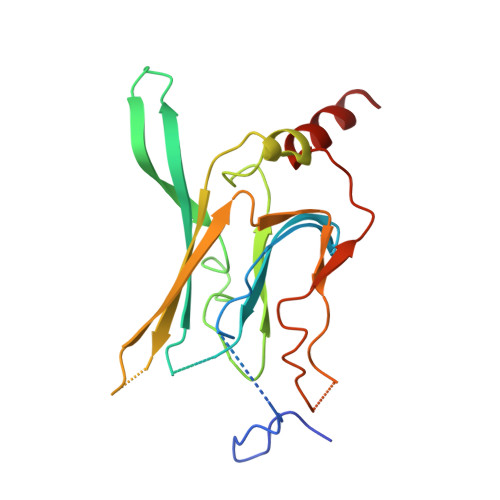
Physical determinants of strong voltage sensitivity of K(+) channel block.
Xu, Y., Shin, H.G., Szep, S., Lu, Z.(2009) Nat Struct Mol Biol 16: 1252-1258
- PubMed: 19915587 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1717
- Primary Citation Related Structures:
3K6N - PubMed Abstract:
Strong voltage sensitivity of inward-rectifier K(+) (Kir) channels has been hypothesized to arise primarily from an intracellular blocker displacing up to five K(+) ions from the wide, intracellular part of the ion conduction pore outwardly across the narrow ion-selectivity filter. The validity of this hypothesis depends on two assumptions: (i) that five ion sites are located intracellular to the filter and (ii) that the blocker can force essentially unidirectional K(+) movement in a pore region generally wider than the combined dimensions of the blocker plus a K(+) ion. Here we present a crystal structure of the cytoplasmic portion of a Kir channel with five ions bound and demonstrate that a constriction near the intracellular end of the pore, acting as a gasket, prevents K(+) ions from bypassing the blocker. This heretofore unrecognized 'gasket' ensures that the blocker can effectively displace K(+) ions across the selectivity filter to generate exceedingly strong voltage sensitivity.
- Department of Physiology, Howard Hughes Medical Institute, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: