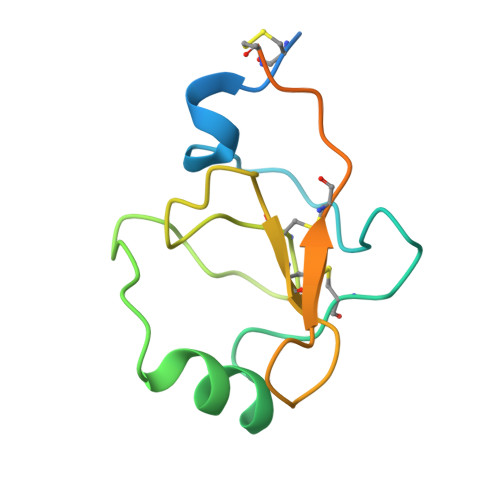
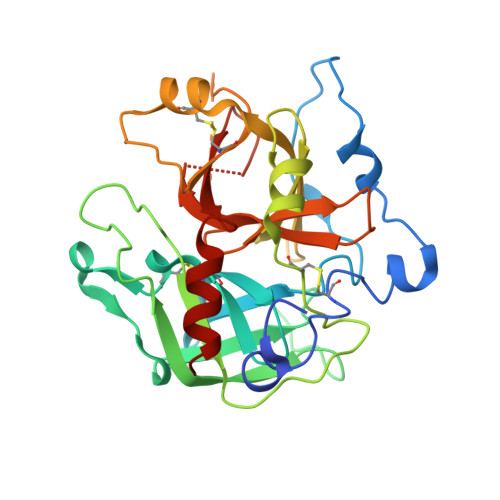
Structural transitions during prothrombin activation: On the importance of fragment 2.
Adams, T.E., Huntington, J.A.(2016) Biochimie 122: 235-242
- PubMed: 26365066 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.biochi.2015.09.013
- Primary Citation Related Structures:
3K65 - PubMed Abstract:
Prothrombin is activated to thrombin by the prothrombinase complex through sequential cleavage at two distinct sites. This occurs at sites of vascular injury in a highly regulated cascade of serine protease and cofactor activation, where activated platelets provide a suitable surface for protease/cofactor/substrate assembly. The precise structural and conformational changes undergone during the transition from prothrombin to thrombin have been studied for decades, and several structures of prothrombin fragments along the activation pathway have been solved. Here we present a new structure analyzed in context of other recent structures and biochemical studies. What emerges is an unexpected mechanism that involves a change in the mode of binding of the F2 domain (fragment 2) on the catalytic domain after cleavage at Arg320, and a subsequent reorientation of the linker between the F2 and catalytic domain to present the Arg271 site for cleavage.
- Cambridge Institute for Medical Research, Department of Haematology, University of Cambridge, Wellcome Trust/MRC Building, Hills Road, Cambridge, CB2 0XY, United Kingdom.
Organizational Affiliation: