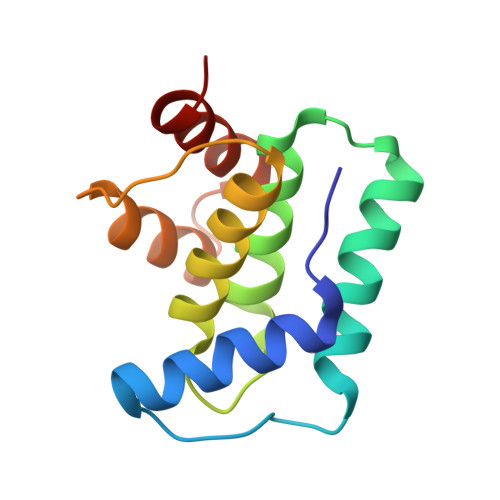
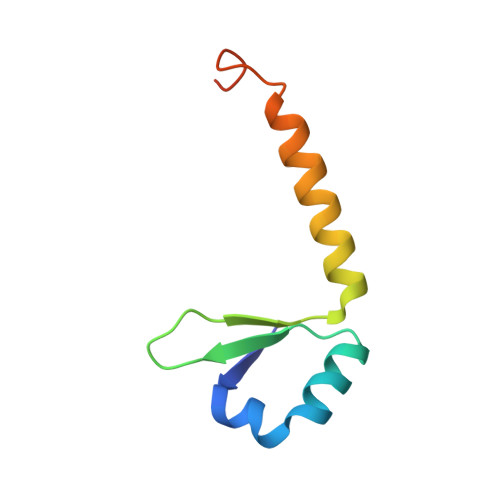
Allostery and intrinsic disorder mediate transcription regulation by conditional cooperativity.
Garcia-Pino, A., Balasubramanian, S., Wyns, L., Gazit, E., De Greve, H., Magnuson, R.D., Charlier, D., van Nuland, N.A., Loris, R.(2010) Cell 142: 101-111
- PubMed: 20603017 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2010.05.039
- Primary Citation Related Structures:
3HRY, 3HS2, 3K33 - PubMed Abstract:
Regulation of the phd/doc toxin-antitoxin operon involves the toxin Doc as co- or derepressor depending on the ratio between Phd and Doc, a phenomenon known as conditional cooperativity. The mechanism underlying this observed behavior is not understood. Here we show that monomeric Doc engages two Phd dimers on two unrelated binding sites. The binding of Doc to the intrinsically disordered C-terminal domain of Phd structures its N-terminal DNA-binding domain, illustrating allosteric coupling between highly disordered and highly unstable domains. This allosteric effect also couples Doc neutralization to the conditional regulation of transcription. In this way, higher levels of Doc tighten repression up to a point where the accumulation of toxin triggers the production of Phd to counteract its action. Our experiments provide the basis for understanding the mechanism of conditional cooperative regulation of transcription typical of toxin-antitoxin modules. This model may be applicable for the regulation of other biological systems.
- Structural Biology Brussels, Vrije Universiteit Brussels, Pleinlaan 2, B-1050 Brussels, Belgium. agarciap@vub.ac.be
Organizational Affiliation: