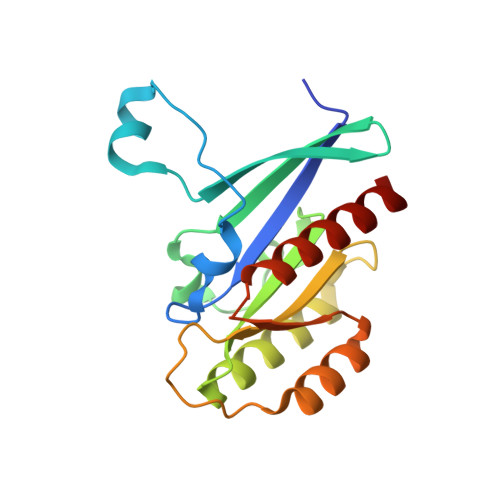
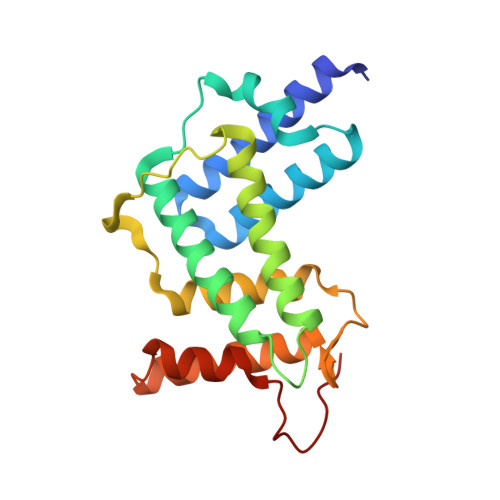
RabGDI displacement by DrrA from Legionella is a consequence of its guanine nucleotide exchange activity.
Schoebel, S., Oesterlin, L.K., Blankenfeldt, W., Goody, R.S., Itzen, A.(2009) Mol Cell 36: 1060-1072
- PubMed: 20064470 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2009.11.014
- Primary Citation Related Structures:
3JZ9, 3JZA - PubMed Abstract:
Prenylated Rab proteins exist in the cytosol as soluble, high-affinity complexes with GDI that need to be disrupted for membrane attachment and targeting of Rab proteins. The Legionella pneumophila protein DrrA displaces GDI from Rab1:GDI complexes, incorporating Rab1 into Legionella-containing vacuoles and activating Rab1 by exchanging GDP for GTP. Here, we present the crystal structure of a complex between the GEF domain of DrrA and Rab1 and a detailed kinetic analysis of this exchange. DrrA efficiently catalyzes nucleotide exchange and mimics the general nucleotide exchange mechanism of mammalian GEFs for Ras-like GTPases. We show that the GEF activity of DrrA is sufficient to displace prenylated Rab1 from the Rab1:GDI complex. Thus, apparent GDI displacement by DrrA is linked directly to nucleotide exchange, suggesting a basic model for GDI displacement and specificity of Rab localization that does not require discrete GDI displacement activity.
- Department of Physical Biochemistry, Max Planck Institute of Molecular Physiology, Dortmund, NRW, Germany.
Organizational Affiliation: