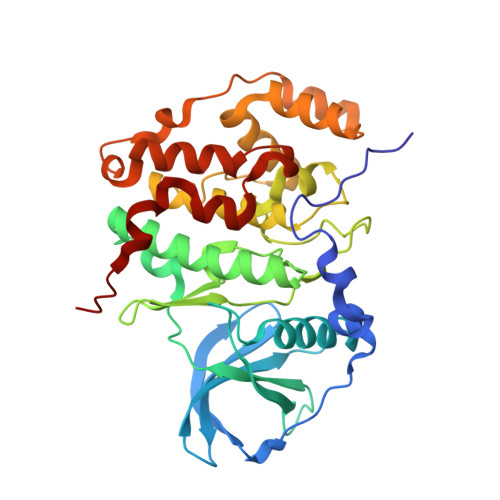
Inclining the purine base binding plane in protein kinase CK2 by exchanging the flanking side-chains generates a preference for ATP as a cosubstrate
Yde, C.W., Ermakova, I., Issinger, O.-G., Niefind, K.(2005) J Mol Biology 347: 399-414
- PubMed: 15740749 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.01.003
- Primary Citation Related Structures:
1LP4, 1LPU, 1LR4, 3JUH - PubMed Abstract:
Protein kinase CK2 (casein kinase 2) is a highly conserved and ubiquitously found eukaryotic serine/threonine kinase that plays a role in various cellular key processes like proliferation, apoptosis and circadian rhythm. One of its prominent biochemical properties is its ability to use GTP as well as ATP as a cosubstrate (dual-cosubstrate specificity). This feature is exceptional among eukaryotic protein kinases, and its biological significance is unknown. We describe here a mutant of the catalytic subunit of protein kinase CK2 (CK2alpha) from Homo sapiens (hsCK2alpha) with a clear and CK2-atypical preference for ATP compared to GTP. This mutant was designed on the basis of several structures of CK2alpha from Zea mays (zmCK2alpha) in complex with various ATP-competitive ligands. A structural overlay revealed the existence of a "purine base binding plane" harbouring the planar moiety of the respective ligand like the purine base of ATP and GTP. This purine base binding plane is sandwiched between the side-chains of Ile66 (Val66 in hsCK2alpha) and Met163, and it adopts a significantly different orientation than in prominent homologues like cAMP-dependent protein kinase (CAPK). By exchanging these two flanking amino acids (Val66Ala, Met163Leu) in hsCK2alpha(1-335), a C-terminally truncated variant of hsCK2alpha, the cosubstrate specificity shifted in the expected direction so that the mutant strongly favours ATP. A structure determination of the mutant in complex with an ATP-analogue confirmed the predicted change of the purine base binding plane orientation. An unexpected but in retrospect plausible consequence of the mutagenesis was, that the helix alpha D region, which is in the direct neighbourhood of the ATP-binding site, has adopted a conformation that is more similar to CAPK and less favourable for binding of GTP. These findings demonstrate that CK2alpha possesses sophisticated structural adaptations in favour of dual-cosubstrate specificity, suggesting that this property could be of biological significance.
- Syddansk Universitet, Institut for Biokemi og Molekylaerbiologi, Campusvej 55, DK-5230 Odense, Denmark.
Organizational Affiliation: