Synthesis, kinase inhibitory potencies, and in vitro antiproliferative evaluation of new pim kinase inhibitors.
Akue-Gedu, R., Rossignol, E., Azzaro, S., Knapp, S., Filippakopoulos, P., Bullock, A.N., Bain, J., Cohen, P., Prudhomme, M., Anizon, F., Moreau, P.(2009) J Med Chem 52: 6369-6381
- PubMed: 19788246 Search on PubMed
- DOI: https://doi.org/10.1021/jm901018f
- Primary Citation Related Structures:
3JPV - PubMed Abstract:
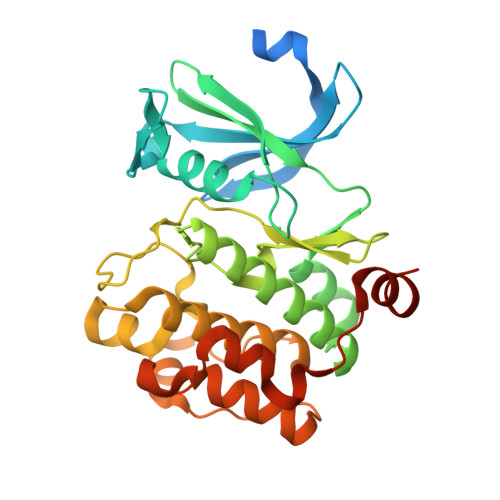
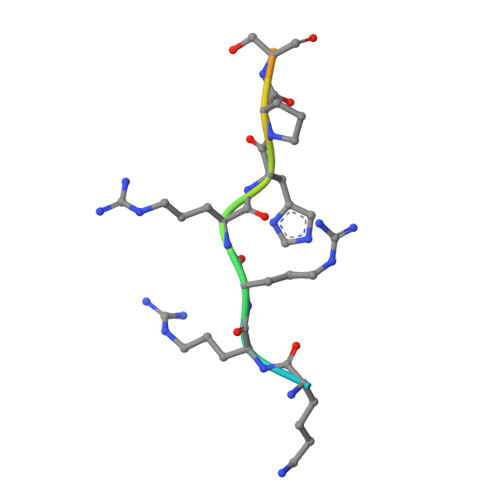
Members of the Pim kinase family have been identified as promising targets for the development of antitumor agents. After a screening of pyrrolo[2,3-a]- and [3,2-a]carbazole derivatives toward 66 protein kinases, we identified pyrrolo[2,3-a]carbazole as a new scaffold to design potent Pim kinase inhibitors. In particular, compound 9 was identified as a low nM selective Pim inhibitor. Additionally, several pyrrolo[2,3-a]carbazole derivatives showed selectivity for Pim-1 and Pim-3 over Pim-2. In vitro antiproliferative activities of 9 and 28, the most potent Pim inhibitors identified, were evaluated toward three human solid cancer cell lines (PA1, PC3, and DU145) and one human fibroblast primary culture, revealing IC50 values in the micromolar range. Finally, the crystal structure of Pim-1 complexed with lead compound 9 was determined. The structure revealed a non-ATP mimetic binding mode with no hydrogen bonds formed with the kinase hinge region and explained the selectivity of pyrrolo[2,3-a]carbazole derivatives for Pim kinases.
- Clermont Université, Université Blaise Pascal, Laboratoire SEESIB, F-63177 Aubière, France.
Organizational Affiliation: