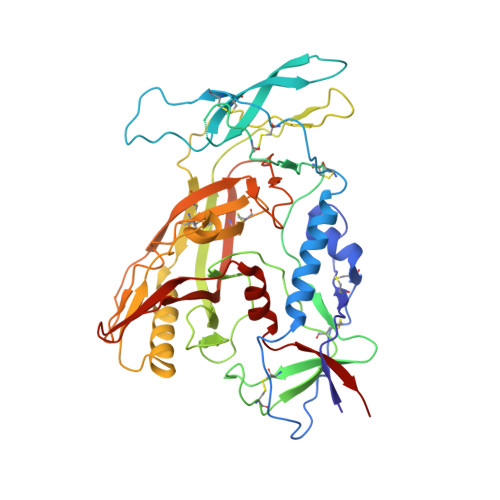
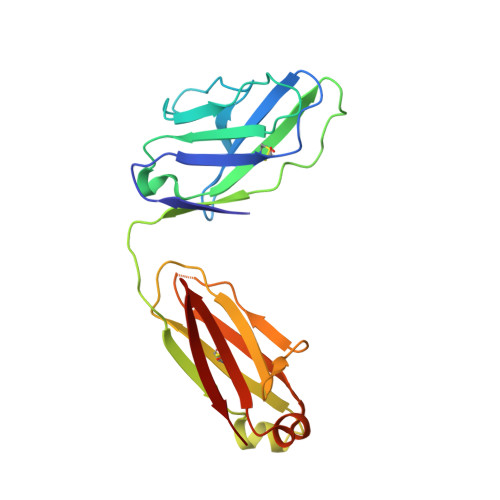
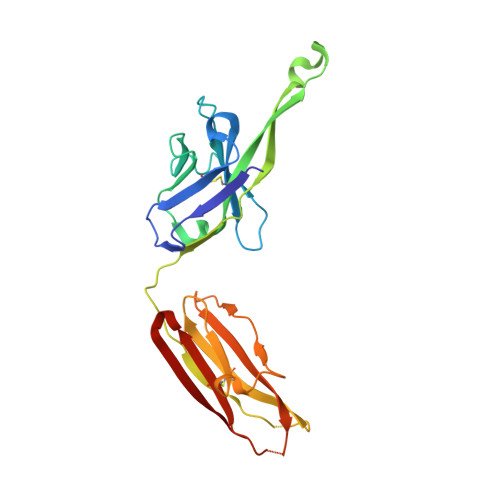
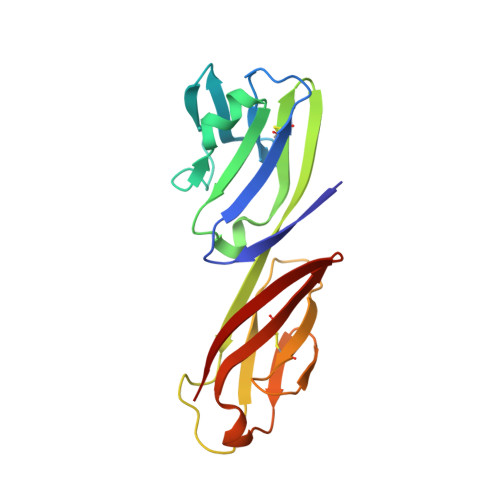
Structure of Simian Immunodeficiency Virus Envelope Spikes Bound with CD4 and Monoclonal Antibody 36D5.
Hu, G., Liu, J., Roux, K.H., Taylor, K.A.(2017) J Virol 91
- PubMed: 28539445 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00134-17
- Primary Citation Related Structures:
3JCB, 3JCC - PubMed Abstract:
The human immunodeficiency virus type 1 (HIV-1)/simian immunodeficiency virus (SIV) envelope spike (Env) mediates viral entry into host cells. The V3 loop of the gp120 component of the Env trimer contributes to the coreceptor binding site and is a target for neutralizing antibodies. We used cryo-electron tomography to visualize the binding of CD4 and the V3 loop monoclonal antibody (MAb) 36D5 to gp120 of the SIV Env trimer. Our results show that 36D5 binds gp120 at the base of the V3 loop and suggest that the antibody exerts its neutralization effect by blocking the coreceptor binding site. The antibody does this without altering the dynamics of the spike motion between closed and open states when CD4 is bound. The interaction between 36D5 and SIV gp120 is similar to the interaction between some broadly neutralizing anti-V3 loop antibodies and HIV-1 gp120. Two conformations of gp120 bound with CD4 are revealed, suggesting an intrinsic dynamic nature of the liganded Env trimer. CD4 binding substantially increases the binding of 36D5 to gp120 in the intact Env trimer, consistent with CD4-induced changes in the conformation of gp120 and the antibody binding site. Binding by MAb 36D5 does not substantially alter the proportions of the two CD4-bound conformations. The position of MAb 36D5 at the V3 base changes little between conformations, indicating that the V3 base serves as a pivot point during the transition between these two states. IMPORTANCE Glycoprotein spikes on the surfaces of SIV and HIV are the sole targets available to the immune system for antibody neutralization. Spikes evade the immune system by a combination of a thick layer of polysaccharide on the surface (the glycan shield) and movement between spike domains that masks the epitope conformation. Using SIV virions whose spikes were "decorated" with the primary cellular receptor (CD4) and an antibody (36D5) at part of the coreceptor binding site, we visualized multiple conformations trapped by the rapid freezing step, which were separated using statistical analysis. Our results show that the CD4-induced conformational dynamics of the spike enhances binding of the antibody.
- Institute of Molecular Biophysics, Florida State University, Tallahassee, Florida, USA.
Organizational Affiliation: