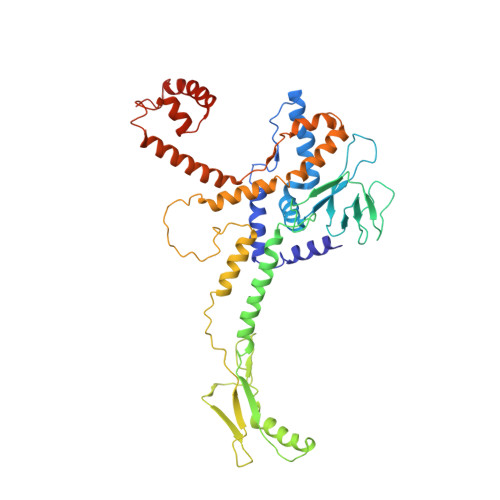
Cryo-EM structure of the bacteriophage T4 portal protein assembly at near-atomic resolution.
Sun, L., Zhang, X., Gao, S., Rao, P.A., Padilla-Sanchez, V., Chen, Z., Sun, S., Xiang, Y., Subramaniam, S., Rao, V.B., Rossmann, M.G.(2015) Nat Commun 6: 7548-7548
- PubMed: 26144253 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8548
- Primary Citation Related Structures:
3JA7 - PubMed Abstract:
The structure and assembly of bacteriophage T4 has been extensively studied. However, the detailed structure of the portal protein remained unknown. Here we report the structure of the bacteriophage T4 portal assembly, gene product 20 (gp20), determined by cryo-electron microscopy (cryo-EM) to 3.6 Å resolution. In addition, analysis of a 10 Å resolution cryo-EM map of an empty prolate T4 head shows how the dodecameric portal assembly interacts with the capsid protein gp23 at the special pentameric vertex. The gp20 structure also verifies that the portal assembly is required for initiating head assembly, for attachment of the packaging motor, and for participation in DNA packaging. Comparison of the Myoviridae T4 portal structure with the known portal structures of φ29, SPP1 and P22, representing Podo- and Siphoviridae, shows that the portal structure probably dates back to a time when self-replicating microorganisms were being established on Earth.
- Department of Biological Sciences, Purdue University, 240S. Martin Jischke Drive, West Lafayette, Indiana 47907-2032, USA.
Organizational Affiliation: