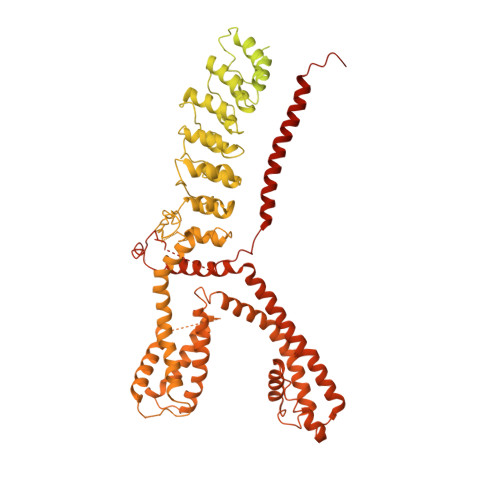
Structure of the TRPA1 ion channel suggests regulatory mechanisms.
Paulsen, C.E., Armache, J.-P., Gao, Y., Cheng, Y., Julius, D.(2015) Nature 520: 511-517
- PubMed: 25855297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature14367
- Primary Citation Related Structures:
3J9P - PubMed Abstract:
The TRPA1 ion channel (also known as the wasabi receptor) is a detector of noxious chemical agents encountered in our environment or produced endogenously during tissue injury or drug metabolism. These include a broad class of electrophiles that activate the channel through covalent protein modification. TRPA1 antagonists hold potential for treating neurogenic inflammatory conditions provoked or exacerbated by irritant exposure. Despite compelling reasons to understand TRPA1 function, structural mechanisms underlying channel regulation remain obscure. Here we use single-particle electron cryo- microscopy to determine the structure of full-length human TRPA1 to ∼4 Å resolution in the presence of pharmacophores, including a potent antagonist. Several unexpected features are revealed, including an extensive coiled-coil assembly domain stabilized by polyphosphate co-factors and a highly integrated nexus that converges on an unpredicted transient receptor potential (TRP)-like allosteric domain. These findings provide new insights into the mechanisms of TRPA1 regulation, and establish a blueprint for structure-based design of analgesic and anti-inflammatory agents.
- Department of Physiology, University of California, San Francisco, California 94158-2517, USA.
Organizational Affiliation: