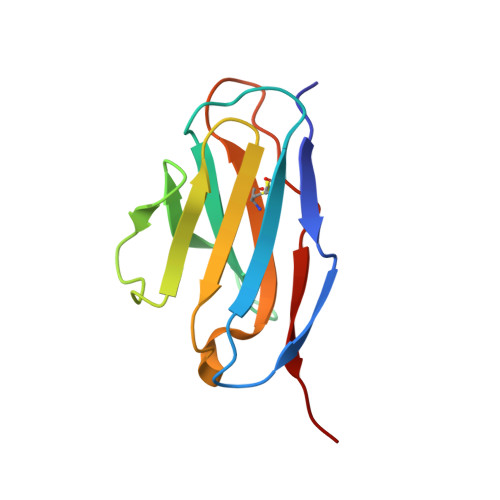
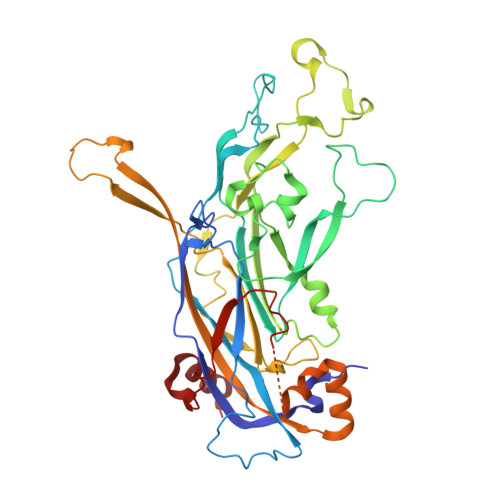
Structural comparison of four different antibodies interacting with human papillomavirus 16 and mechanisms of neutralization.
Guan, J., Bywaters, S.M., Brendle, S.A., Lee, H., Ashley, R.E., Makhov, A.M., Conway, J.F., Christensen, N.D., Hafenstein, S.(2015) Virology 483: 253-263
- PubMed: 25996608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.virol.2015.04.016
- Primary Citation Related Structures:
3J8V, 3J8W, 3J8Z - PubMed Abstract:
Cryo-electron microscopy (cryo-EM) was used to solve the structures of human papillomavirus type 16 (HPV16) complexed with fragments of antibody (Fab) from three different neutralizing monoclonals (mAbs): H16.1A, H16.14J, and H263.A2. The structure-function analysis revealed predominantly monovalent binding of each Fab with capsid interactions that involved multiple loops from symmetry related copies of the major capsid protein. The residues identified in each Fab-virus interface map to a conformational groove on the surface of the capsomer. In addition to the known involvement of the FG and HI loops, the DE loop was also found to constitute the core of each epitope. Surprisingly, the epitope mapping also identified minor contributions by EF and BC loops. Complementary immunological assays included mAb and Fab neutralization. The specific binding characteristics of mAbs correlated with different neutralizing behaviors in pre- and post-attachment neutralization assays.
- Department of Medicine, The Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA 17033 USA.
Organizational Affiliation: