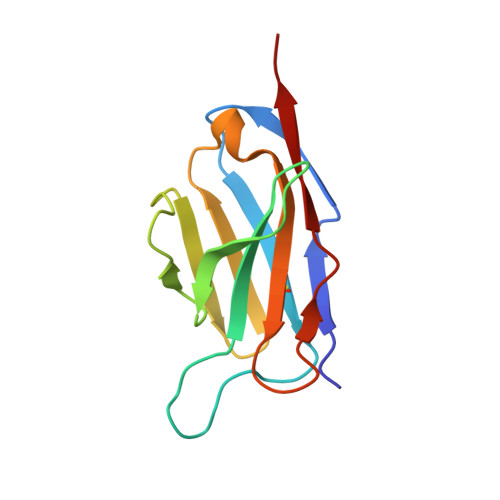
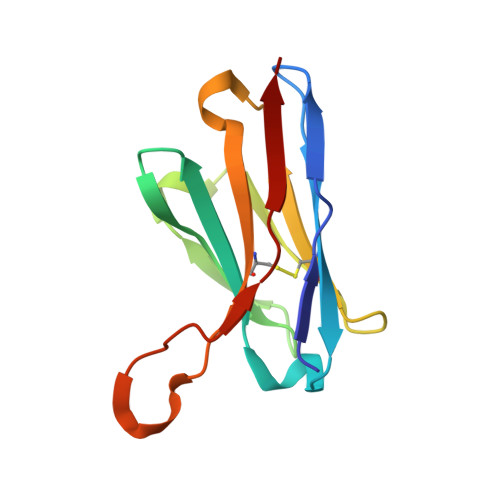
A cryo-electron microscopy study identifies the complete H16.V5 epitope and reveals global conformational changes initiated by binding of the neutralizing antibody fragment.
Lee, H., Brendle, S.A., Bywaters, S.M., Guan, J., Ashley, R.E., Yoder, J.D., Makhov, A.M., Conway, J.F., Christensen, N.D., Hafenstein, S.(2015) J Virol 89: 1428-1438
- PubMed: 25392224
- DOI: https://doi.org/10.1128/JVI.02898-14
- Primary Citation Related Structures:
3J7E, 3J7G - PubMed Abstract:
Human papillomavirus 16 (HPV16) is a worldwide health threat and an etiologic agent of cervical cancer. To understand the antigenic properties of HPV16, we pursued a structural study to elucidate HPV capsids and antibody interactions. The cryo-electron microscopy (cryo-EM) structures of a mature HPV16 particle and an altered capsid particle were solved individually and as complexes with fragment of antibody (Fab) from the neutralizing antibody H16.V5. Fitted crystal structures provided a pseudoatomic model of the virus-Fab complex, which identified a precise footprint of H16.V5, including previously unrecognized residues. The altered-capsid-Fab complex map showed that binding of the Fab induced significant conformational changes that were not seen in the altered-capsid structure alone. These changes included more ordered surface loops, consolidated so-called "invading-arm" structures, and tighter intercapsomeric connections at the capsid floor. The H16.V5 Fab preferentially bound hexavalent capsomers likely with a stabilizing effect that directly correlated with the number of bound Fabs. Additional cryo-EM reconstructions of the virus-Fab complex for different incubation times and structural analysis provide a model for a hyperstabilization of the capsomer by H16.V5 Fab and showed that the Fab distinguishes subtle differences between antigenic sites. Our analysis of the cryo-EM reconstructions of the HPV16 capsids and virus-Fab complexes has identified the entire HPV.V5 conformational epitope and demonstrated a detailed neutralization mechanism of this clinically important monoclonal antibody against HPV16. The Fab bound and ordered the apical loops of HPV16. This conformational change was transmitted to the lower region of the capsomer, resulting in enhanced intercapsomeric interactions evidenced by the more ordered capsid floor and "invading-arm" structures. This study advances the understanding of the neutralization mechanism used by H16.V5.
- Department of Medicine, The Pennsylvania State University College of Medicine, Hershey, Pennsylvania, USA.
Organizational Affiliation: