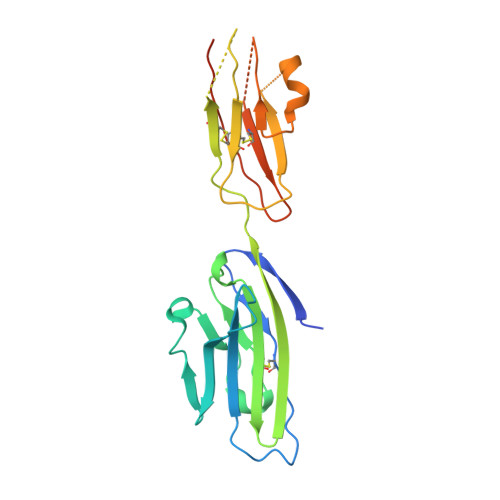
Kinetic and structural analysis of coxsackievirus b3 receptor interactions and formation of the a-particle.
Organtini, L.J., Makhov, A.M., Conway, J.F., Hafenstein, S., Carson, S.D.(2014) J Virol 88: 5755-5765
- PubMed: 24623425 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00299-14
- Primary Citation Related Structures:
3J6L, 3J6M, 3J6N, 3J6O - PubMed Abstract:
The coxsackievirus and adenovirus receptor (CAR) has been identified as the cellular receptor for group B coxsackieviruses, including serotype 3 (CVB3). CAR mediates infection by binding to CVB3 and catalyzing conformational changes in the virus that result in formation of the altered, noninfectious A-particle. Kinetic analyses show that the apparent first-order rate constant for the inactivation of CVB3 by soluble CAR (sCAR) at physiological temperatures varies nonlinearly with sCAR concentration. Cryo-electron microscopy (cryo-EM) reconstruction of the CVB3-CAR complex resulted in a 9.0-Å resolution map that was interpreted with the four available crystal structures of CAR, providing a consensus footprint for the receptor binding site. The analysis of the cryo-EM structure identifies important virus-receptor interactions that are conserved across picornavirus species. These conserved interactions map to variable antigenic sites or structurally conserved regions, suggesting a combination of evolutionary mechanisms for receptor site preservation. The CAR-catalyzed A-particle structure was solved to a 6.6-Å resolution and shows significant rearrangement of internal features and symmetric interactions with the RNA genome. This report presents new information about receptor use by picornaviruses and highlights the importance of attaining at least an ∼9-Å resolution for the interpretation of cryo-EM complex maps. The analysis of receptor binding elucidates two complementary mechanisms for preservation of the low-affinity (initial) interaction of the receptor and defines the kinetics of receptor-catalyzed conformational change to the A-particle.
- Department of Medicine, The Pennsylvania State University College of Medicine, Hershey, Pennsylvania, USA.
Organizational Affiliation: