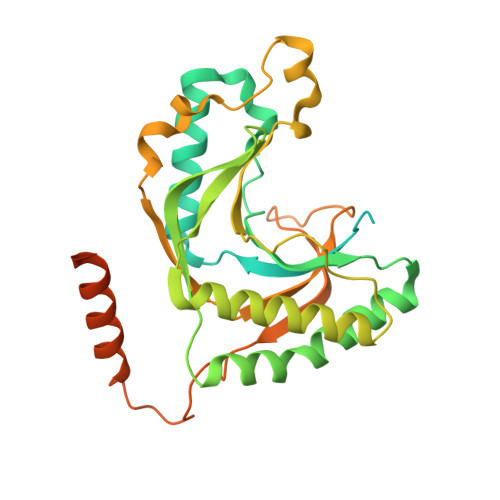
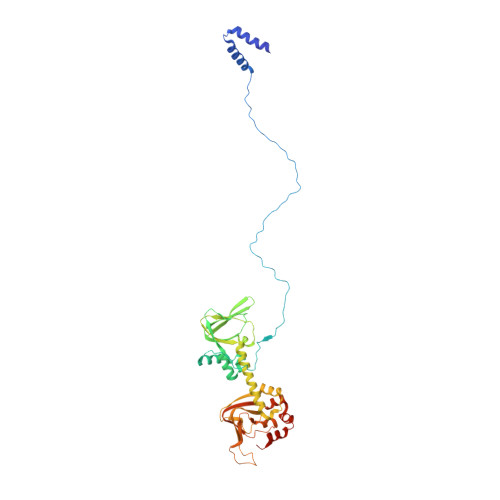
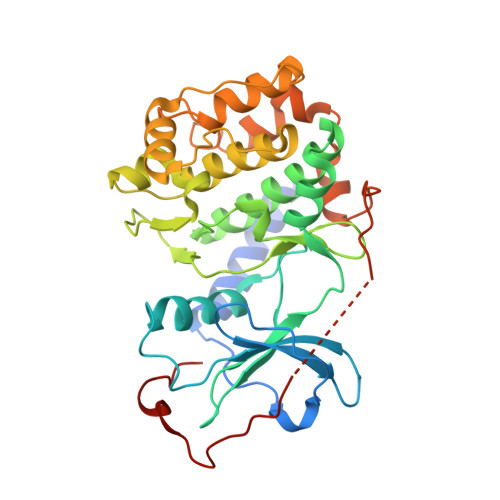
Intrinsic disorder within an AKAP-protein kinase A complex guides local substrate phosphorylation.
Smith, F.D., Reichow, S.L., Esseltine, J.L., Shi, D., Langeberg, L.K., Scott, J.D., Gonen, T.(2013) Elife 2: e01319-e01319
- PubMed: 24192038
- DOI: https://doi.org/10.7554/eLife.01319
- Primary Citation Related Structures:
3J4Q, 3J4R - PubMed Abstract:
Anchoring proteins sequester kinases with their substrates to locally disseminate intracellular signals and avert indiscriminate transmission of these responses throughout the cell. Mechanistic understanding of this process is hampered by limited structural information on these macromolecular complexes. A-kinase anchoring proteins (AKAPs) spatially constrain phosphorylation by cAMP-dependent protein kinases (PKA). Electron microscopy and three-dimensional reconstructions of type-II PKA-AKAP18γ complexes reveal hetero-pentameric assemblies that adopt a range of flexible tripartite configurations. Intrinsically disordered regions within each PKA regulatory subunit impart the molecular plasticity that affords an ∼16 nanometer radius of motion to the associated catalytic subunits. Manipulating flexibility within the PKA holoenzyme augmented basal and cAMP responsive phosphorylation of AKAP-associated substrates. Cell-based analyses suggest that the catalytic subunit remains within type-II PKA-AKAP18γ complexes upon cAMP elevation. We propose that the dynamic movement of kinase sub-structures, in concert with the static AKAP-regulatory subunit interface, generates a solid-state signaling microenvironment for substrate phosphorylation. DOI: http://dx.doi.org/10.7554/eLife.01319.001.
- Department of Pharmacology, Howard Hughes Medical Institute, University of Washington, Seattle, United States.
Organizational Affiliation: