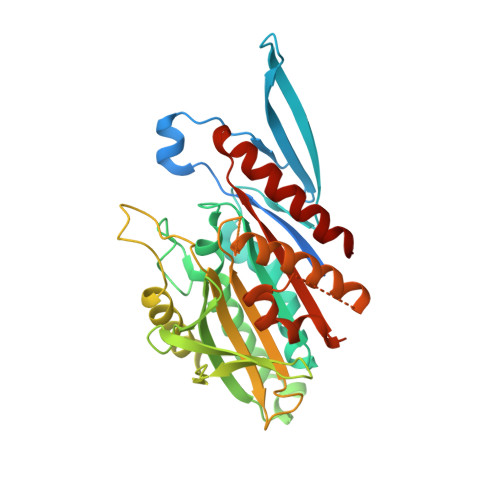
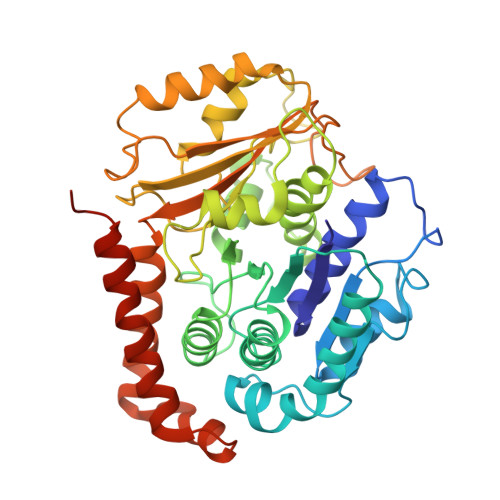
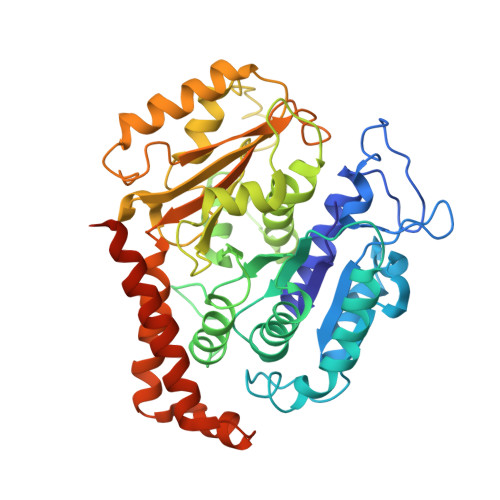
Structural model for tubulin recognition and deformation by Kinesin-13 microtubule depolymerases.
Asenjo, A.B., Chatterjee, C., Tan, D., Depaoli, V., Rice, W.J., Diaz-Avalos, R., Silvestry, M., Sosa, H.(2013) Cell Rep 3: 759-768
- PubMed: 23434508 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2013.01.030
- Primary Citation Related Structures:
3J2U - PubMed Abstract:
To elucidate the structural basis of the mechanism of microtubule depolymerization by kinesin-13s, we analyzed complexes of tubulin and the Drosophila melanogaster kinesin-13 KLP10A by electron microscopy (EM) and fluorescence polarization microscopy. We report a nanometer-resolution (1.1 nm) cryo-EM three-dimensional structure of the KLP10A head domain (KLP10AHD) bound to curved tubulin. We found that binding of KLP10AHD induces a distinct tubulin configuration with displacement (shear) between tubulin subunits in addition to curvature. In this configuration, the kinesin-binding site differs from that in straight tubulin, providing an explanation for the distinct interaction modes of kinesin-13s with the microtubule lattice or its ends. The KLP10AHD-tubulin interface comprises three areas of interaction, suggesting a crossbow-type tubulin-bending mechanism. These areas include the kinesin-13 family conserved KVD residues, and as predicted from the crossbow model, mutating these residues changes the orientation and mobility of KLP10AHDs interacting with the microtubule.
- Department of Physiology and Biophysics, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: