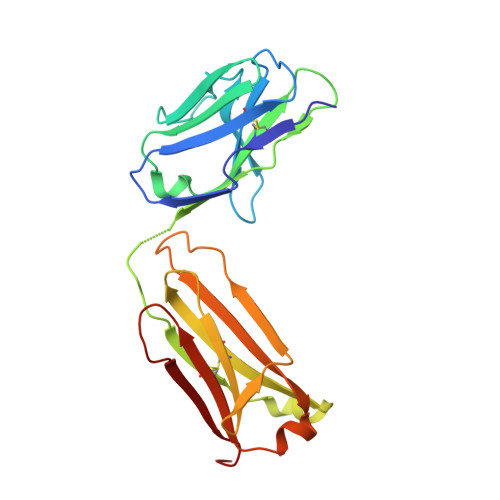
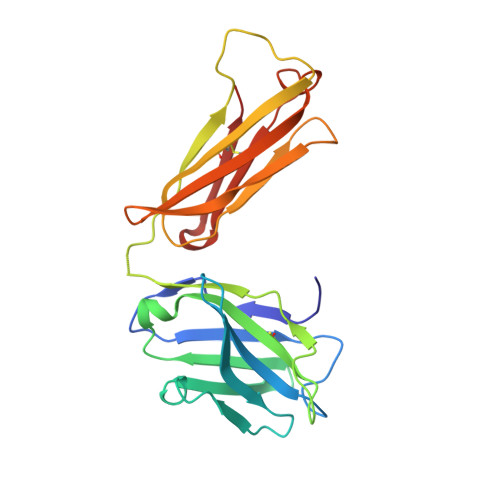
Structure of adeno-associated virus-2 in complex with neutralizing monoclonal antibody A20.
McCraw, D.M., O'Donnell, J.K., Taylor, K.A., Stagg, S.M., Chapman, M.S.(2012) Virology 431: 40-49
- PubMed: 22682774 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.virol.2012.05.004
- Primary Citation Related Structures:
3J1S - PubMed Abstract:
The use of adeno-associated virus (AAV) as a gene therapy vector is limited by the host neutralizing immune response. The cryo-electron microscopy (EM) structure at 8.5Å resolution is determined for a complex of AAV-2 with the Fab' fragment of monoclonal antibody (MAb) A20, the most extensively characterized AAV MAb. The binding footprint is determined through fitting the cryo-EM reconstruction with a homology model following sequencing of the variable domain, and provides a structural basis for integrating diverse prior epitope mappings. The footprint extends from the previously implicated plateau to the side of the spike, and into the conserved canyon, covering a larger area than anticipated. Comparison with structures of binding and non-binding serotypes indicates that recognition depends on a combination of subtle serotype-specific features. Separation of the neutralizing epitope from the heparan sulfate cell attachment site encourages attempts to develop immune-resistant vectors that can still bind to target cells.
- Department of Biochemistry and Molecular Biology, School of Medicine, Mail code L224, Oregon Health & Science University, 3181 S.W. Sam Jackson Park Road, Portland, OR 97239-3098, USA.
Organizational Affiliation: