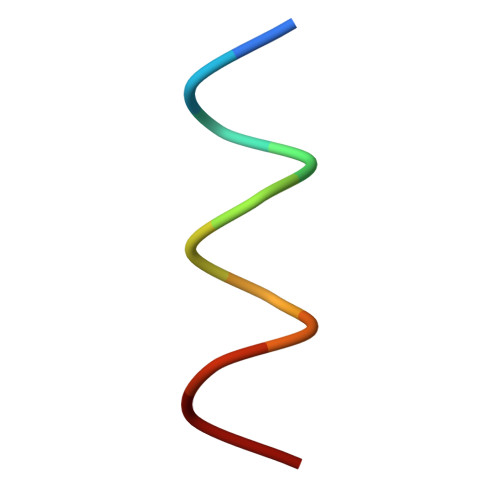Catching a virus in the act of RNA release: a novel poliovirus uncoating intermediate characterized by cryo-electron microscopy.
Levy, H.C., Bostina, M., Filman, D.J., Hogle, J.M.(2010) J Virol 84: 4426-4441
- PubMed: 20181687 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02393-09
- Primary Citation Related Structures:
3IYB, 3IYC - PubMed Abstract:
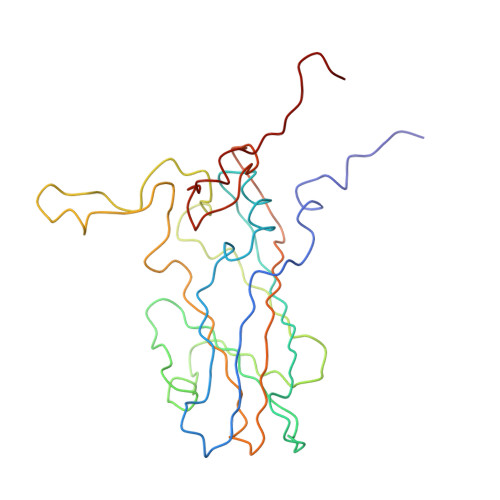
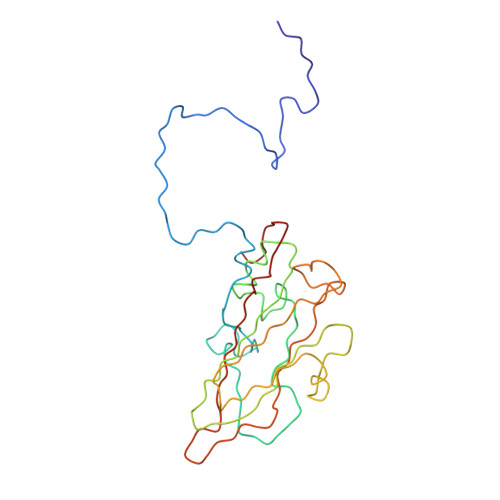
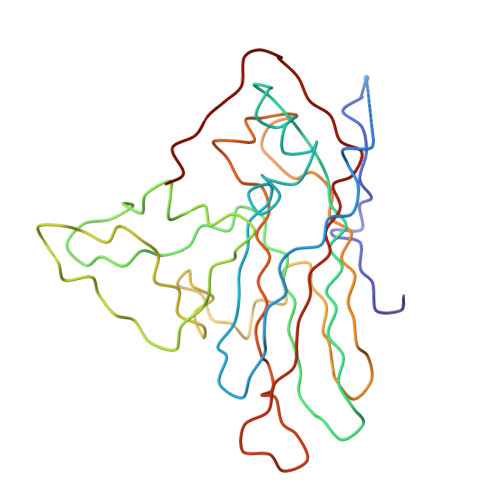
Poliovirus infection requires that the particle undergo a series of conformational transitions that lead to cell entry and genome release. In an effort to understand the conformational changes associated with the release of the RNA genome, we have used cryo-electron microscopy to characterize the structure of the 80S "empty" particles of poliovirus that are thought to represent the final product of the cell entry pathway. Using two-dimensional classification methods, we show that preparations of 80S particles contain at least two structures, which might represent snapshots from a continuous series of conformers. Using three-dimensional reconstruction methods, we have solved the structure of two distinct forms at subnanometric resolution, and we have built and refined pseudoatomic models into the reconstructions. The reconstructions and the derived models demonstrate that the two structural forms are both slightly expanded, resulting in partial disruption of interprotomer interfaces near their particle 2-fold axes, which may represent the site where RNA is released. The models demonstrate that each of the two 80S structures has undergone a unique set of movements of the capsid proteins, associated with rearrangement of flexible loops and amino-terminal extensions that participate in contacts between protomers, between pentamers, and with the viral RNA.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts, USA.
Organizational Affiliation: