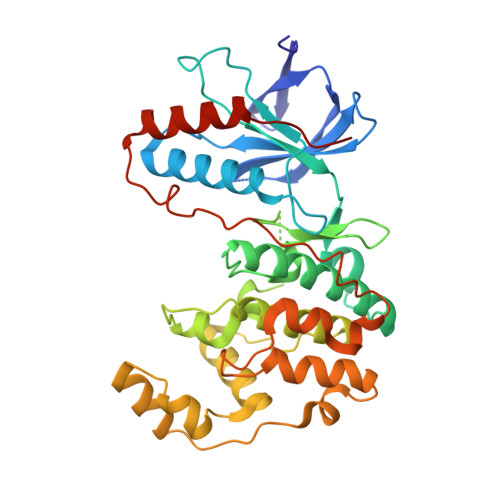
High-Throughput Screening To Identify Inhibitors Which Stabilize Inactive Kinase Conformations in p38alpha
Simard, J.R., Gruetter, C., Pawar, V., Aust, B., Wolf, A., Rabiller, M., Wulfert, S., Robubi, A., Kluter, S., Ottmann, C., Rauh, D.(2009) J Am Chem Soc 131: 18478-18488
- PubMed: 19950957 Search on PubMed
- DOI: https://doi.org/10.1021/ja907795q
- Primary Citation Related Structures:
3IW5, 3IW6, 3IW7, 3IW8 - PubMed Abstract:
Small molecule kinase inhibitors are an attractive means to modulate kinase activities in medicinal chemistry and chemical biology research. In the physiological setting of a cell, kinase function is orchestrated by a plethora of regulatory processes involving the structural transition of kinases between inactive and enzymatically competent conformations and vice versa. The development of novel kinase inhibitors is mainly fostered by high-throughput screening initiatives where the small molecule perturbation of the phosphorylation reaction is measured to identify inhibitors. Such setups require enzymatically active kinase preparations and present a risk of solely identifying classical ATP-competitive Type I inhibitors. Here we report the high-throughput screening of a library of approximately 35000 small organic molecules with an assay system that utilizes enzymatically inactive human p38alpha MAP kinase to detect stabilizers of the pharmacologically more desirable DFG-out conformation. We used protein X-ray crystallography to characterize the binding mode of hit compounds and reveal structural features which explain how these ligands stabilize and/or induce the DFG-out conformation. Lastly, we show that although some of the hit compounds were confirmed by protein X-ray crystallography, they were not detected in classic phosphorylation assays, thus validating the unique sensitivity of the assay system used in this study and highlighting the potential of screening with inactive kinase preparations.
- Chemical Genomics Centre of the Max Planck Society, Otto-Hahn-Strasse 15, D-44227 Dortmund, Germany.
Organizational Affiliation: