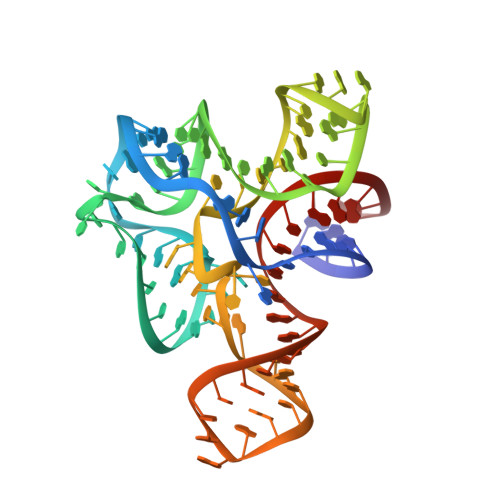
Free state conformational sampling of the SAM-I riboswitch aptamer domain.
Stoddard, C.D., Montange, R.K., Hennelly, S.P., Rambo, R.P., Sanbonmatsu, K.Y., Batey, R.T.(2010) Structure 18: 787-797
- PubMed: 20637415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.04.006
- Primary Citation Related Structures:
3IQN, 3IQP, 3IQR - PubMed Abstract:
Riboswitches are highly structured elements residing in the 5' untranslated region of messenger RNAs that specifically bind cellular metabolites to alter gene expression. While there are many structures of ligand-bound riboswitches that reveal details of bimolecular recognition, their unliganded structures remain poorly characterized. Characterizing the molecular details of the unliganded state is crucial for understanding the riboswitch's mechanism of action because it is this state that actively interrogates the cellular environment and helps direct the regulatory outcome. To develop a detailed description of the ligand-free form of an S-adenosylmethionine binding riboswitch at the local and global levels, we have employed a series of biochemical, biophysical, and computational methods. Our data reveal that the ligand binding domain adopts an ensemble of states that minimizes the energy barrier between the free and bound states to establish an efficient decision making branchpoint in the regulatory process.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, UCB 215, Boulder, CO 80309-0215, USA.
Organizational Affiliation: