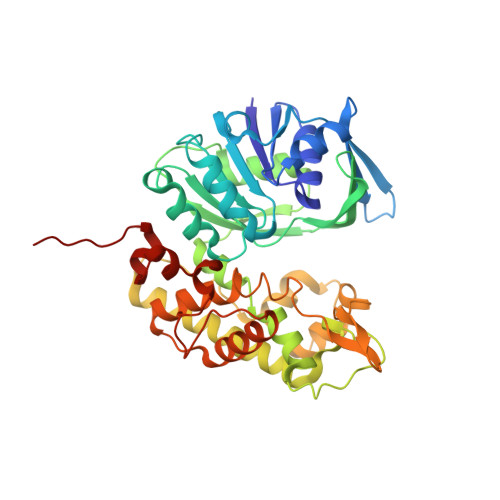
Insights into the mechanism of ligand binding to octopine dehydrogenase from Pecten maximus by NMR and crystallography
Smits, S.H.J., Meyer, T., Mueller, A., van Os, N., Stoldt, M., Willbold, D., Schmitt, L., Grieshaber, M.K.(2010) PLoS One 5: e12312-e12312
- PubMed: 20808820 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0012312
- Primary Citation Related Structures:
3IQD - PubMed Abstract:
Octopine dehydrogenase (OcDH) from the adductor muscle of the great scallop, Pecten maximus, catalyzes the NADH dependent, reductive condensation of L-arginine and pyruvate to octopine, NAD(+), and water during escape swimming and/or subsequent recovery. The structure of OcDH was recently solved and a reaction mechanism was proposed which implied an ordered binding of NADH, L-arginine and finally pyruvate. Here, the order of substrate binding as well as the underlying conformational changes were investigated by NMR confirming the model derived from the crystal structures. Furthermore, the crystal structure of the OcDH/NADH/agmatine complex was determined which suggests a key role of the side chain of L-arginine in protein cataylsis. Thus, the order of substrate binding to OcDH as well as the molecular signals involved in octopine formation can now be described in molecular detail.
- Institute of Biochemistry, Heinrich-Heine-University, Duesseldorf, Germany. sander.smits@uni-duesseldorf.de
Organizational Affiliation: