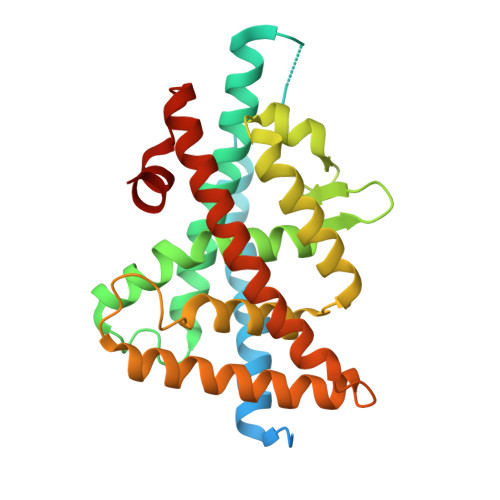
X-ray structures of the LXRalpha LBD in its homodimeric form and implications for heterodimer signaling.
Fradera, X., Vu, D., Nimz, O., Skene, R., Hosfield, D., Wynands, R., Cooke, A.J., Haunso, A., King, A., Bennett, D.J., McGuire, R., Uitdehaag, J.C.(2010) J Mol Biology 399: 120-132
- PubMed: 20382159 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.04.005
- Primary Citation Related Structures:
3IPQ, 3IPS, 3IPU - PubMed Abstract:
Liver X receptors (LXRs) are nuclear receptors that are central regulators of cholesterol homeostasis, and synthetic LXR agonists have shown promise as promoters of reverse cholesterol transport and anti-inflammatory agents. Here, we present three X-ray structures of three different agonists bound to the ligand binding domain of LXRalpha. These compounds are GW3965, F(3)methylAA, and a benzisoxazole urea, and we show that these diverse chemical scaffolds address common structural themes, leading to high binding affinity for LXR. Our structures show the LXRalpha ligand binding domain in its homodimeric form, an arrangement previously thought to be stereochemically difficult. A comparison with existing structures of the LXRbeta homodimer and LXRalpha:RXR (retinoid X receptor) heterodimers explains differences in dimer affinity and leads us to propose a model for allosteric activation in nuclear receptor dimers, in which an unactivated RXR partner provides an inhibitory tail wrap to the cofactor binding pocket of LXR.
- Department of Chemistry, Merck Research Labs, Newhouse, ML1 5SH, Scotland, UK.
Organizational Affiliation: