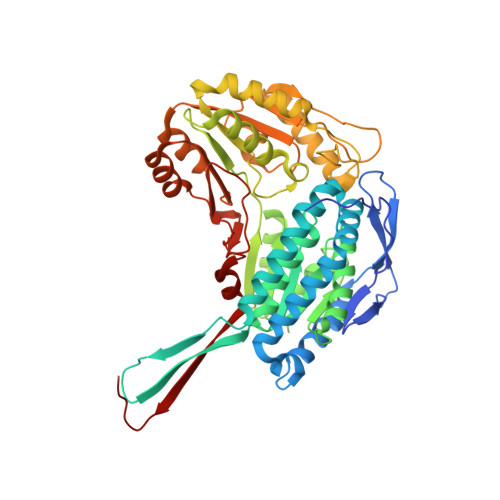
Alda-1 is an agonist and chemical chaperone for the common human aldehyde dehydrogenase 2 variant.
Perez-Miller, S., Younus, H., Vanam, R., Chen, C.H., Mochly-Rosen, D., Hurley, T.D.(2010) Nat Struct Mol Biol 17: 159-164
- PubMed: 20062057 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1737
- Primary Citation Related Structures:
3INJ, 3INL - PubMed Abstract:
In approximately one billion people, a point mutation inactivates a key detoxifying enzyme, aldehyde dehydrogenase (ALDH2). This mitochondrial enzyme metabolizes toxic biogenic and environmental aldehydes, including the endogenously produced 4-hydroxynonenal (4HNE) and the environmental pollutant acrolein, and also bioactivates nitroglycerin. ALDH2 is best known, however, for its role in ethanol metabolism. The accumulation of acetaldehyde following the consumption of even a single alcoholic beverage leads to the Asian alcohol-induced flushing syndrome in ALDH2*2 homozygotes. The ALDH2*2 allele is semidominant, and heterozygotic individuals show a similar but less severe phenotype. We recently identified a small molecule, Alda-1, that activates wild-type ALDH2 and restores near-wild-type activity to ALDH2*2. The structures of Alda-1 bound to ALDH2 and ALDH2*2 reveal how Alda-1 activates the wild-type enzyme and how it restores the activity of ALDH2*2 by acting as a structural chaperone.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana, USA.
Organizational Affiliation: