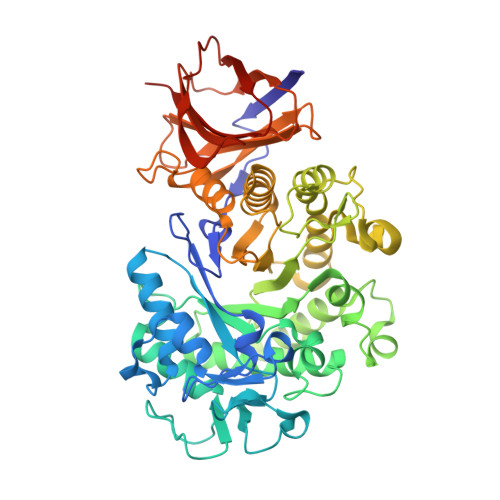
Tertiary structure and characterization of a glycoside hydrolase family 44 endoglucanase from Clostridium acetobutylicum.
Warner, C.D., Hoy, J.A., Shilling, T.C., Linnen, M.J., Ginder, N.D., Ford, C.F., Honzatko, R.B., Reilly, P.J.(2010) Appl Environ Microbiol 76: 338-346
- PubMed: 19915043
- DOI: https://doi.org/10.1128/AEM.02026-09
- Primary Citation Related Structures:
3IK2 - PubMed Abstract:
A gene encoding a glycoside hydrolase family 44 (GH44) protein from Clostridium acetobutylicum ATCC 824 was synthesized and transformed into Escherichia coli. The previously uncharacterized protein was expressed with a C-terminal His tag and purified by nickel-nitrilotriacetic acid affinity chromatography. Crystallization and X-ray diffraction to a 2.2-A resolution revealed a triose phosphate isomerase (TIM) barrel-like structure with additional Greek key and beta-sandwich folds, similar to other GH44 crystal structures. The enzyme hydrolyzes cellotetraose and larger cellooligosaccharides, yielding an unbalanced product distribution, including some glucose. It attacks carboxymethylcellulose and xylan at approximately the same rates. Its activity on carboxymethylcellulose is much higher than that of the isolated C. acetobutylicum cellulosome. It also extensively converts lichenan to oligosaccharides of intermediate size and attacks Avicel to a limited extent. The enzyme has an optimal temperature in a 10-min assay of 55 degrees C and an optimal pH of 5.0.
- Department of Chemical and Biological Engineering, 2114 Sweeney Hall, Iowa State University, Ames, IA 50011-2230, USA.
Organizational Affiliation: