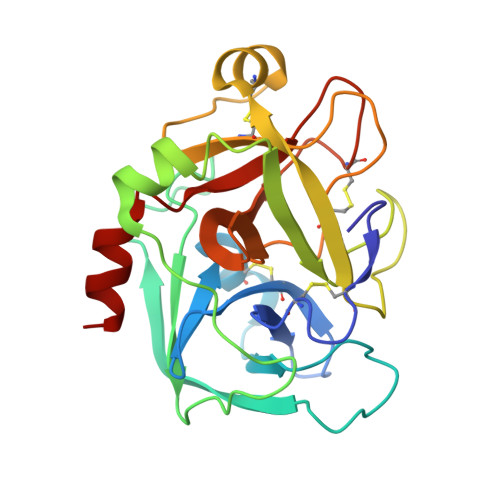
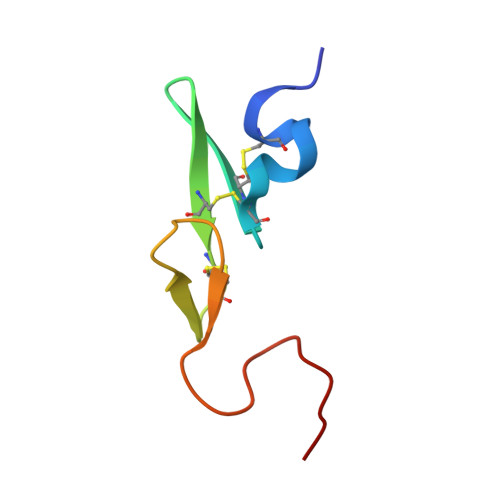
Design, synthesis, and SAR of cis-1,2-diaminocyclohexane derivatives as potent factor Xa inhibitors. Part II: exploration of 6-6 fused rings as alternative S1 moieties.
Yoshikawa, K., Kobayashi, S., Nakamoto, Y., Haginoya, N., Komoriya, S., Yoshino, T., Nagata, T., Mochizuki, A., Watanabe, K., Suzuki, M., Kanno, H., Ohta, T.(2009) Bioorg Med Chem 17: 8221-8233
- PubMed: 19900814 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2009.10.024
- Primary Citation Related Structures:
3IIT - PubMed Abstract:
A series of cis-1,2-diaminocyclohexane derivatives possessing a 6-6 fused ring for the S1 moiety were synthesized as novel factor Xa (fXa) inhibitors. The synthesis, structure-activity relationship (SAR), and physicochemical properties are reported herein, together with the discovery of compound 45c, which has potent anti-fXa activity, good physicochemical properties and pharmacokinetic (PK) profiles, including a reduced negative food effect.
- R&D Division, Daiichi Sankyo Co, Ltd, 1-16-13, Kita-Kasai, Tokyo 134-8630, Japan. yoshikawa.kenji.t6@daiichisankyo.co.jp
Organizational Affiliation: