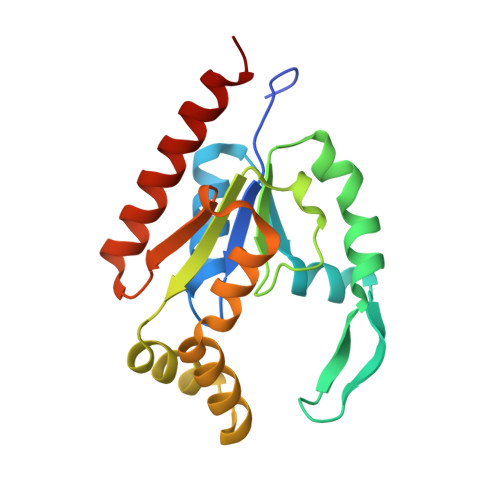
hCINAP is an atypical mammalian nuclear adenylate kinase with an ATPase motif: Structural and functional studies.
Drakou, C.E., Malekkou, A., Hayes, J.M., Lederer, C.W., Leonidas, D.D., Oikonomakos, N.G., Lamond, A.I., Santama, N., Zographos, S.E.(2012) Proteins 80: 206-220
- PubMed: 22038794 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.23186
- Primary Citation Related Structures:
3IIJ, 3IIK, 3IIL, 3IIM - PubMed Abstract:
Human coilin interacting nuclear ATPase protein (hCINAP) directly interacts with coilin, a marker protein of Cajal Bodies (CBs), nuclear organelles involved in the maturation of small nuclear ribonucleoproteins UsnRNPs and snoRNPs. hCINAP has previously been designated as an adenylate kinase (AK6), but is very atypical as it exhibits unusually broad substrate specificity, structural features characteristic of ATPase/GTPase proteins (Walker motifs A and B) and also intrinsic ATPase activity. Despite its intriguing structure, unique properties and cellular localization, the enzymatic mechanism and biological function of hCINAP have remained poorly characterized. Here, we offer the first high-resolution structure of hCINAP in complex with the substrate ADP (and dADP), the structure of hCINAP with a sulfate ion bound at the AMP binding site, and the structure of the ternary complex hCINAP-Mg(2+) ADP-Pi. Induced fit docking calculations are used to predict the structure of the hCINAP-Mg(2+) ATP-AMP ternary complex. Structural analysis suggested a functional role for His79 in the Walker B motif. Kinetic analysis of mutant hCINAP-H79G indicates that His79 affects both AK and ATPase catalytic efficiency and induces homodimer formation. Finally, we show that in vivo expression of hCINAP-H79G in human cells is toxic and drastically deregulates the number and appearance of CBs in the cell nucleus. Our findings suggest that hCINAP may not simply regulate nucleotide homeostasis, but may have broader functionality, including control of CB assembly and disassembly in the nucleus of human cells.
- Institute of Organic and Pharmaceutical Chemistry, National Hellenic Research Foundation, Athens 11635, Greece.
Organizational Affiliation: