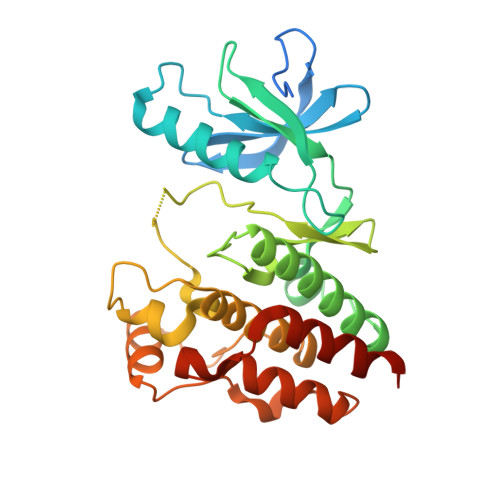
Non-hinge-binding pyrazolo[1,5-a]pyrimidines as potent B-Raf kinase inhibitors.
Berger, D.M., Torres, N., Dutia, M., Powell, D., Ciszewski, G., Gopalsamy, A., Levin, J.I., Kim, K.H., Xu, W., Wilhelm, J., Hu, Y., Collins, K., Feldberg, L., Kim, S., Frommer, E., Wojciechowicz, D., Mallon, R.(2009) Bioorg Med Chem Lett 19: 6519-6523
- PubMed: 19864136 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.10.049
- Primary Citation Related Structures:
3II5 - PubMed Abstract:
As part of our research effort to discover B-Raf kinase inhibitors, we prepared a series of C-3 substituted N-(3-(pyrazolo[1,5-a]pyrimidin-7-yl)phenyl)-3-(trifluoromethyl)benzamides. X-ray crystallography studies revealed that one of the more potent inhibitors (10n) bound to B-Raf kinase without forming a hinge-binding hydrogen bond. With basic amine residues appended to C-3 aryl residues, cellular activity and solubility were enhanced over previously described compounds of this class.
- Chemical Sciences, Wyeth Research, Pearl River, NY 10965, USA. bergerd@wyeth.com
Organizational Affiliation: