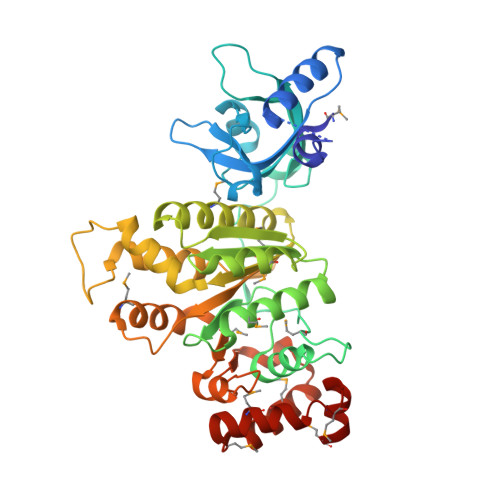
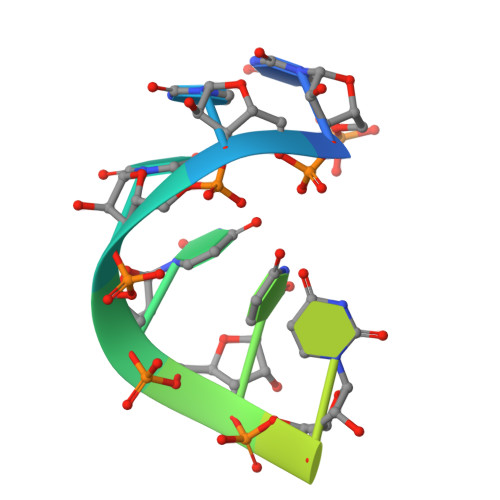
Running in reverse: the structural basis for translocation polarity in hexameric helicases.
Thomsen, N.D., Berger, J.M.(2009) Cell 139: 523-534
- PubMed: 19879839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2009.08.043
- Primary Citation Related Structures:
3ICE - PubMed Abstract:
Hexameric helicases couple ATP hydrolysis to processive separation of nucleic acid duplexes, a process critical for gene expression, DNA replication, and repair. All hexameric helicases fall into two families with opposing translocation polarities: the 3'-->5' AAA+ and 5'-->3' RecA-like enzymes. To understand how a RecA-like hexameric helicase engages and translocates along substrate, we determined the structure of the E. coli Rho transcription termination factor bound to RNA and nucleotide. Interior nucleic acid-binding elements spiral around six bases of RNA in a manner unexpectedly reminiscent of an AAA+ helicase, the papillomavirus E1 protein. Four distinct ATP-binding states, representing potential catalytic intermediates, are coupled to RNA positioning through a complex allosteric network. Comparative studies with E1 suggest that RecA and AAA+ hexameric helicases use different portions of their chemomechanical cycle for translocating nucleic acid and track in opposite directions by reversing the firing order of ATPase sites around the hexameric ring. For a video summary of this article, see the PaperFlick file with the Supplemental Data available online.
- Department of Molecular and Cell Biology, Quantitative Biosciences Institute, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: