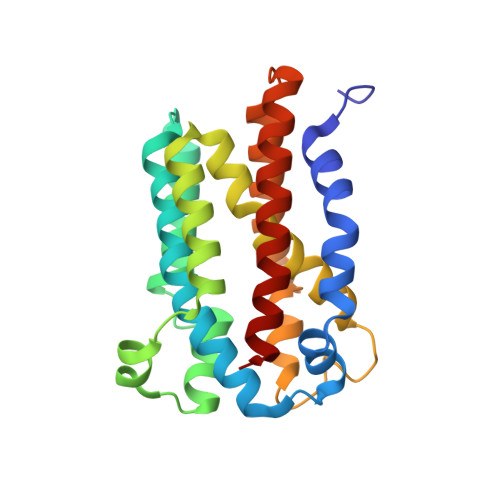
Structural and mutational analysis of TenA protein (HP1287) from the Helicobacter pylori thiamin salvage pathway - evidence of a different substrate specificity.
Barison, N., Cendron, L., Trento, A., Angelini, A., Zanotti, G.(2009) FEBS J 276: 6227-6235
- PubMed: 19780837 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2009.07326.x
- Primary Citation Related Structures:
3IBX - PubMed Abstract:
HP1287 (tenA) from Helicobacter pylori is included among the genes that play a relevant role in bacterium colonization and persistence. The gene has been cloned and its product, protein TenA, has been expressed and purified. The crystal structures of the wild-type protein and the mutant F47Y have been determined at resolutions of 2.7 and 2.4 A, respectively. The molecular model, a homotetramer with 222 symmetry, shows that the H. pylori TenA structure belongs to the thiaminase II class of proteins. These enzymes were recently found to be involved in a salvage pathway for the synthesis of the thiamin precursor hydroxypyrimidine, which constitutes a building block in thiamin biosynthesis, in particular in bacteria living in the soil. By contrast, enzymatic measurements on TenA from H. pylori indicate that the activity on the putative substrate 4-amino-5-aminomethyl-2-methylpyrimidine is very modest. Moreover, in the present study, we demonstrate that the mutation at residue 47, a position where a phenylalanine occurs in all the strains of H. pylori sequenced to date, is not sufficient to explain the very low catalytic activity toward the expected substrate. As a result of differences in the colonization environment of H. pylori as well as the TenA structural and catalytic peculiar features, we suggest a possible pivotal role for the H. pylori enzyme in the thiamin biosynthetic route, which is in agreement with the relevance of this protein in the stomach colonization process.
- Department of Biological Chemistry, University of Padua, Italy.
Organizational Affiliation: