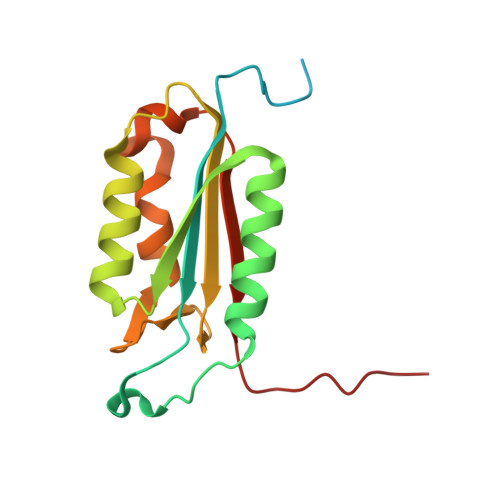
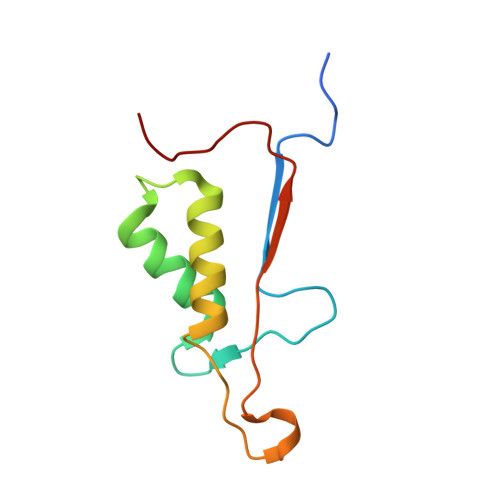
Conformational similarity in the activation of caspase-3 and -7 revealed by the unliganded and inhibited structures of caspase-7.
Agniswamy, J., Fang, B., Weber, I.T.(2009) Apoptosis 14: 1135-1144
- PubMed: 19655253 Search on PubMed
- DOI: https://doi.org/10.1007/s10495-009-0388-9
- Primary Citation Related Structures:
3IBC, 3IBF - PubMed Abstract:
Caspase-mediated apoptosis has important roles in normal cell differentiation and aging and in many diseases including cancer, neuromuscular disorders and neurodegenerative diseases. Therefore, modulation of caspase activity and conformational states is of therapeutic importance. We report crystal structures of a new unliganded conformation of caspase-7 and the inhibited caspase-7 with the tetrapeptide Ac-YVAD-Cho. Different conformational states and mechanisms for substrate recognition have been proposed based on unliganded structures of the redundant apoptotic executioner caspase-3 and -7. The current study shows that the executioner caspase-3 and -7 have similar conformations for the unliganded active site as well as the inhibitor-bound active site. The new unliganded caspase-7 structure exhibits the tyrosine flipping mechanism in which the Tyr230 has rotated to block entry to the S2 binding site similar to the active site conformation of unliganded caspase-3. The inhibited structure of caspase-7/YVAD shows that the P4 Tyr binds the S4 region specific to polar residues at the expense of a main chain hydrogen bond between the P4 amide and carbonyl oxygen of caspase-7 Gln 276, which is similar to the caspase-3 complex. This new knowledge of the structures and conformational states of unliganded and inhibited caspases will be important for the design of drugs to modulate caspase activity and apoptosis.
- Department of Biology, Georgia State University, Atlanta, GA 30303, USA.
Organizational Affiliation: