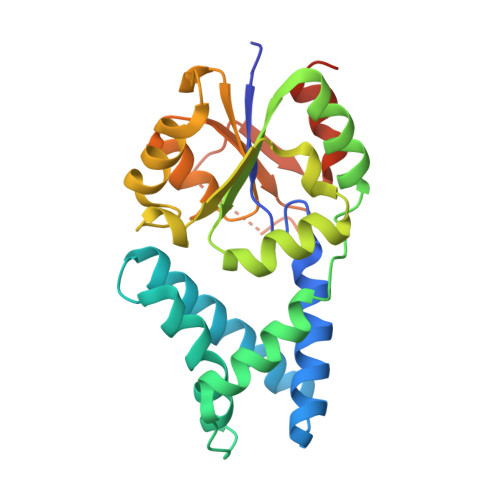
The crystal structure of the orthorhombic form of the putative HAD-hydrolase YfnB from Bacillus subtilis bound to magnesium reveals interdomain movement
Bonanno, J.B., Dickey, M., Bain, K.T., Tang, B.K., Romero, R., Sauder, J.M., Burley, S.K., Almo, S.C.To be published.