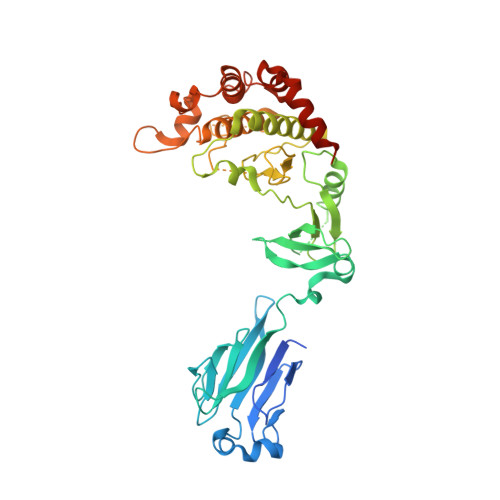
Structure and activation mechanism of the CHK2 DNA damage checkpoint kinase.
Cai, Z., Chehab, N.H., Pavletich, N.P.(2009) Mol Cell 35: 818-829
- PubMed: 19782031 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2009.09.007
- Primary Citation Related Structures:
3I6U, 3I6W - PubMed Abstract:
The CHK2 protein kinase is an important transducer of DNA damage checkpoint signals, and its mutation contributes to hereditary and sporadic cancer. CHK2 activation is triggered by the phosphorylation of Thr68 by the DNA damage-activated ATM kinase. This leads to transient CHK2 dimerization, in part through intermolecular phosphoThr68-FHA domain interactions. Dimerization promotes kinase activation through activation-loop autophosphorylation, but the mechanism of this process has not been clear. The dimeric crystal structure of CHK2, described here, in conjunction with biochemical and mutational data reveals that productive CHK2 dimerization additionally involves intermolecular FHA-kinase domain and FHA-FHA interactions. Ile157, mutated in the Li-Fraumeni cancer-predisposition syndrome, plays a central role in the FHA-kinase domain interface, explaining the lack of dimerization and autophosphorylation of this mutant. In the dimer, the kinase active sites face each other in close proximity, indicating that dimerization may also serve to optimally position the kinase active sites for efficient activation loop transphosphorylation.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10021, USA.
Organizational Affiliation: