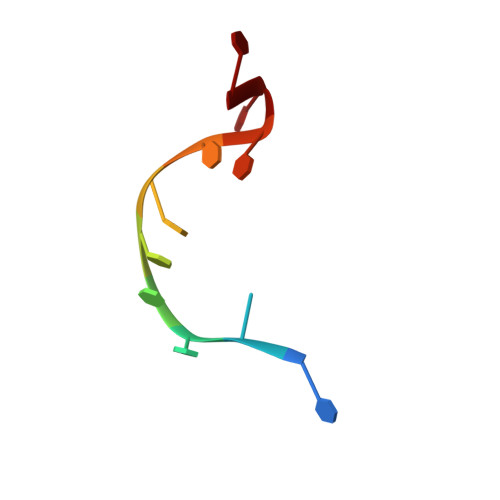
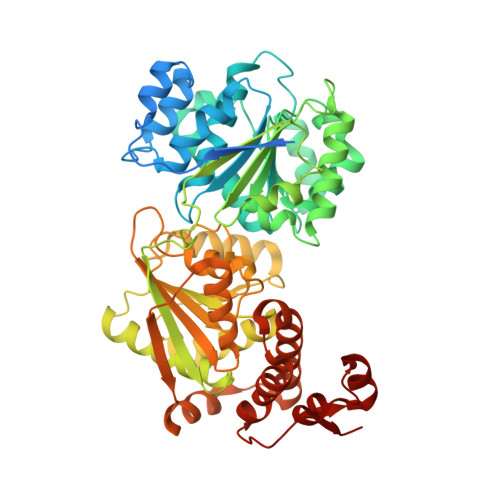
Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA
Del Campo, M., Lambowitz, A.M.(2009) Mol Cell 35: 598-609
- PubMed: 19748356 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2009.07.032
- Primary Citation Related Structures:
3I5X, 3I5Y, 3I61, 3I62 - PubMed Abstract:
The yeast DEAD box protein Mss116p is a general RNA chaperone that functions in mitochondrial group I and II intron splicing, translational activation, and RNA end processing. Here we determined high-resolution X-ray crystal structures of Mss116p complexed with an RNA oligonucleotide and ATP analogs AMP-PNP, ADP-BeF(3)(-), or ADP-AlF(4)(-). The structures show the entire helicase core acting together with a functionally important C-terminal extension. In all structures, the helicase core is in a closed conformation with a wedge alpha helix bending RNA 3' of the central bound nucleotides, as in previous DEAD box protein structures. Notably, Mss116p's C-terminal extension also bends RNA 5' of the central nucleotides, resulting in RNA crimping. Despite reported functional differences, we observe few structural changes in ternary complexes with different ATP analogs. The structures constrain models of DEAD box protein function and reveal a strand separation mechanism in which a protein uses two wedges to act as a molecular crimper.
- Institute for Cellular and Molecular Biology, University of Texas at Austin, 78712, USA.
Organizational Affiliation: