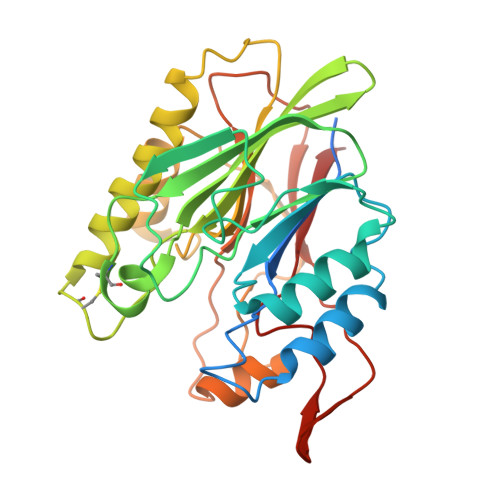
Structure and biological functions of beta toxin from Staphylococcus aureus: Role of the hydrophobic beta hairpin in virulence
Huseby, M., Shi, K., Kruse, A.C., Digre, J., Mengistu, F., Bohach, G.A., Schlievert, P.S., Ohlendorf, D.H., Earhart, C.A.To be published.