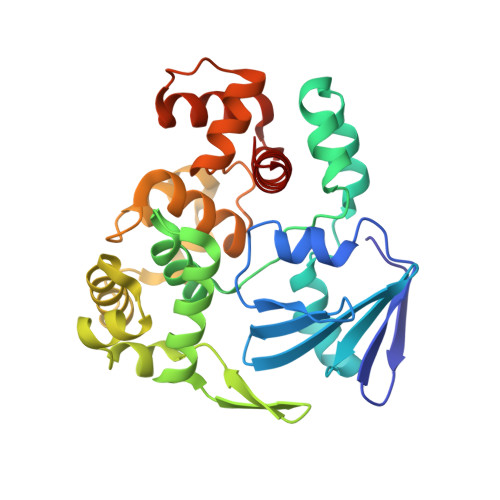
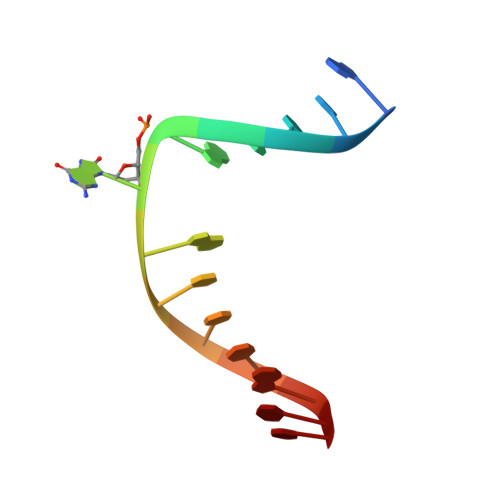
Structural basis for the lack of opposite base specificity of Clostridium acetobutylicum 8-oxoguanine DNA glycosylase.
Faucher, F., Wallace, S.S., Doublie, S.(2009) DNA Repair (Amst) 8: 1283-1289
- PubMed: 19747886 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.dnarep.2009.08.002
- Primary Citation Related Structures:
3I0W, 3I0X - PubMed Abstract:
7,8-Dihydro-8-oxoguanine (8-oxoG) is the major oxidative product of guanine and the most prevalent base lesion observed in DNA molecules. Because 8-oxoG has the capability to form a Hoogsteen pair with adenine (8-oxoG:A) in addition to a normal Watson-Crick pair with cytosine (8-oxoG:C), this lesion can lead to a G:C-->T:A transversion after replication. However, 8-oxoG is recognized and excised by the 8-oxoguanine DNA glycosylase (Ogg) of the base excision repair pathway. Members of the Ogg1 family usually display a strong preference for a C opposite the lesion. In contrast, the atypical Ogg1 from Clostridium actetobutylicum (CacOgg) can excise 8-oxoG when paired with either one of the four bases, albeit with a preference for C and A. Here we describe the first high-resolution crystal structures of CacOgg in complex with duplex DNA containing the 8-oxoG lesion paired to cytosine and to adenine. A structural comparison with human OGG1 provides a rationale for the lack of opposite base specificity displayed by the bacterial Ogg.
- Department of Microbiology and Molecular Genetics, The Markey Center for Molecular Genetics, University of Vermont, Stafford Hall, 95 Carrigan Drive, Burlington, VT 05405-0068, USA.
Organizational Affiliation: