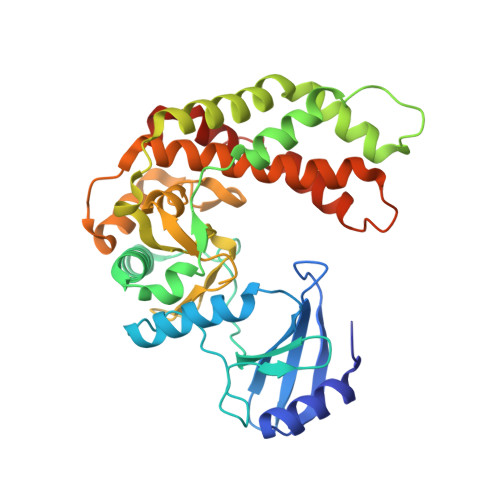
Structure of the antibiotic resistance factor spectinomycin phosphotransferase from Legionella pneumophila.
Fong, D.H., Lemke, C.T., Hwang, J., Xiong, B., Berghuis, A.M.(2010) J Biological Chem 285: 9545-9555
- PubMed: 20089863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.038364
- Primary Citation Related Structures:
3I0O, 3I0Q, 3I1A - PubMed Abstract:
Aminoglycoside phosphotransferases (APHs) constitute a diverse group of enzymes that are often the underlying cause of aminoglycoside resistance in the clinical setting. Several APHs have been extensively characterized, including the elucidation of the three-dimensional structure of two APH(3') isozymes and an APH(2'') enzyme. Although many APHs are plasmid-encoded and are capable of inactivating numerous 2-deoxystreptmaine aminoglycosides with multiple regiospecificity, APH(9)-Ia, isolated from Legionella pneumophila, is an unusual enzyme among the APH family for its chromosomal origin and its specificity for a single non-2-deoxystreptamine aminoglycoside substrate, spectinomycin. We describe here the crystal structures of APH(9)-Ia in its apo form, its binary complex with the nucleotide, AMP, and its ternary complex bound with ADP and spectinomycin. The structures reveal that APH(9)-Ia adopts the bilobal protein kinase-fold, analogous to the APH(3') and APH(2'') enzymes. However, APH(9)-Ia differs significantly from the other two types of APH enzymes in its substrate binding area and that it undergoes a conformation change upon ligand binding. Moreover, kinetic assay experiments indicate that APH(9)-Ia has stringent substrate specificity as it is unable to phosphorylate substrates of choline kinase or methylthioribose kinase despite high structural resemblance. The crystal structures of APH(9)-Ia demonstrate and expand our understanding of the diversity of the APH family, which in turn will facilitate the development of new antibiotics and inhibitors.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6.
Organizational Affiliation: