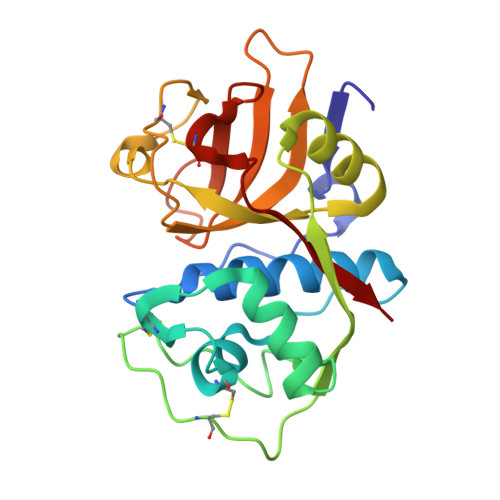
Identification and optimization of inhibitors of trypanosomal cysteine proteases: cruzain, rhodesain, and TbCatB.
Mott, B.T., Ferreira, R.S., Simeonov, A., Jadhav, A., Ang, K.K., Leister, W., Shen, M., Silveira, J.T., Doyle, P.S., Arkin, M.R., McKerrow, J.H., Inglese, J., Austin, C.P., Thomas, C.J., Shoichet, B.K., Maloney, D.J.(2010) J Med Chem 53: 52-60
- PubMed: 19908842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm901069a
- Primary Citation Related Structures:
3I06 - PubMed Abstract:
Trypanosoma cruzi and Trypanosoma brucei are parasites that cause Chagas' disease and African sleeping sickness, respectively. Both parasites rely on essential cysteine proteases for survival: cruzain for T. cruzi and TbCatB/rhodesain for T. brucei. A recent quantitative high-throughput screen of cruzain identified triazine nitriles, which are known inhibitors of other cysteine proteases, as reversible inhibitors of the enzyme. Structural modifications detailed herein, including core scaffold modification from triazine to purine, improved the in vitro potency against both cruzain and rhodesain by 350-fold, while also gaining activity against T. brucei parasites. Selected compounds were screened against a panel of human cysteine and serine proteases to determine selectivity, and a cocrystal was obtained of our most potent analogue bound to cruzain.
- NIH Chemical Genomics Center, National Human Genome Research Institute, National Institutes of Health, 9800 Medical Center Drive, MSC 3370 Bethesda, Maryland 20892-3370, USA.
Organizational Affiliation: