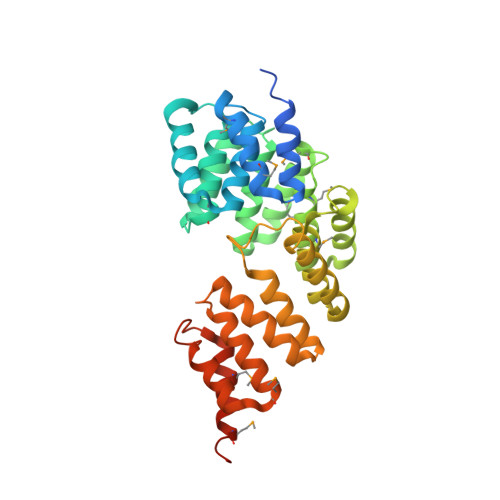
Insights into anaphase promoting complex TPR subdomain assembly from a CDC26-APC6 structure.
Wang, J., Dye, B.T., Rajashankar, K.R., Kurinov, I., Schulman, B.A.(2009) Nat Struct Mol Biol 16: 987-989
- PubMed: 19668213 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1645
- Primary Citation Related Structures:
3HYM - PubMed Abstract:
The multisubunit anaphase promoting complex (APC) is an essential cell-cycle regulator. Although CDC26 is known to have a role in APC assembly, its molecular function has remained unclear. Biophysical, structural and genetic studies presented here reveal that CDC26 stabilizes the structure of APC6, a core TPR protein required for APC integrity. Notably, CDC26-APC6 association involves an intermolecular TPR mimic composed of one helix from each protein.
- Departments of Structural Biology and Genetics/Tumor Cell Biology, St. Jude Children's Research Hospital, Memphis, Tennessee, USA.
Organizational Affiliation: