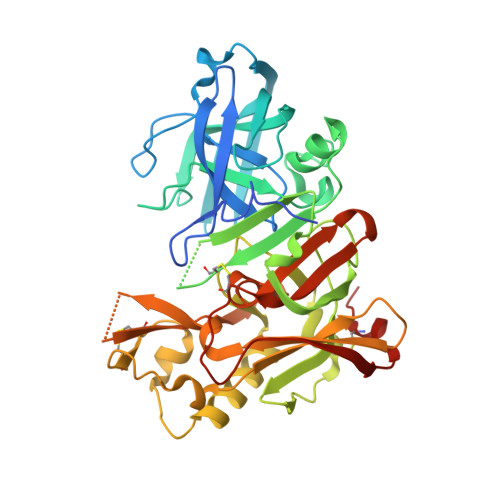
Fragment-Based Discovery of BACE1 Inhibitors Using Functional Assays
Godemann, R., Madden, J., Kramer, J., Smith, M.A., Fritz, U., Hesterkamp, T., Barker, J., Hoeppner, S., Hallett, D., Cesura, A., Ebneth, A., Kemp, J.(2009) Biochemistry 48: 10743-10751
- PubMed: 19799414 Search on PubMed
- DOI: https://doi.org/10.1021/bi901061a
- Primary Citation Related Structures:
3HVG, 3HW1 - PubMed Abstract:
Novel nonpeptidic inhibitors of beta-secretase (BACE1) have been discovered by employing a fragment-based biochemical screening approach. A diverse library of 20000 low-molecular weight compounds were screened and yielded 26 novel hits that were confirmed by biochemical and surface plasmon resonance secondary assays. We describe here fragment inhibitors cocrystallized with BACE1 in a flap open and flap closed conformation as determined by X-ray crystallography.
- Evotec AG, Schnackenburgallee 114, 22525 Hamburg, Germany. robert.godemann@evotec.com
Organizational Affiliation: