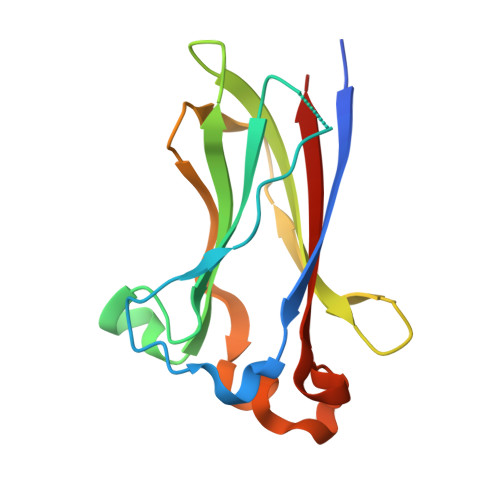
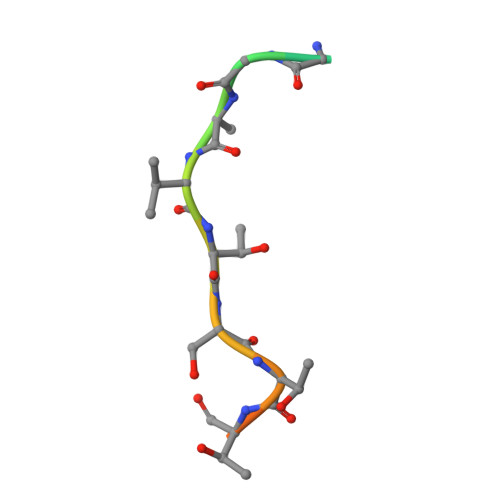
Structures of SPOP-substrate complexes: insights into molecular architectures of BTB-Cul3 ubiquitin ligases.
Zhuang, M., Calabrese, M.F., Liu, J., Waddell, M.B., Nourse, A., Hammel, M., Miller, D.J., Walden, H., Duda, D.M., Seyedin, S.N., Hoggard, T., Harper, J.W., White, K.P., Schulman, B.A.(2009) Mol Cell 36: 39-50
- PubMed: 19818708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2009.09.022
- Primary Citation Related Structures:
3HQH, 3HQI, 3HQL, 3HQM, 3HSV, 3HTM, 3HU6, 3HVE, 3IVQ, 3IVV - PubMed Abstract:
In the largest E3 ligase subfamily, Cul3 binds a BTB domain, and an associated protein-interaction domain such as MATH recruits substrates for ubiquitination. Here, we present biochemical and structural analyses of the MATH-BTB protein, SPOP. We define a SPOP-binding consensus (SBC) and determine structures revealing recognition of SBCs from the phosphatase Puc, the transcriptional regulator Ci, and the chromatin component MacroH2A. We identify a dimeric SPOP-Cul3 assembly involving a conserved helical structure C-terminal of BTB domains, which we call "3-box" due to its facilitating Cul3 binding and its resemblance to F-/SOCS-boxes in other cullin-based E3s. Structural flexibility between the substrate-binding MATH and Cul3-binding BTB/3-box domains potentially allows a SPOP dimer to engage multiple SBCs found within a single substrate, such as Puc. These studies provide a molecular understanding of how MATH-BTB proteins recruit substrates to Cul3 and how their dimerization and conformational variability may facilitate avid interactions with diverse substrates.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN 38105, USA.
Organizational Affiliation: