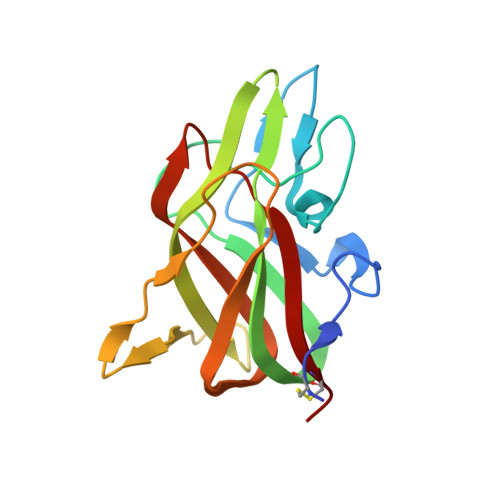
Trp2313-His2315 of factor VIII C2 domain is involved in membrane binding: structure of a complex between the C2 domain and an inhibitor of membrane binding.
Liu, Z., Lin, L., Yuan, C., Nicolaes, G.A., Chen, L., Meehan, E.J., Furie, B., Furie, B., Huang, M.(2010) J Biological Chem 285: 8824-8829
- PubMed: 20089867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.080168
- Primary Citation Related Structures:
3HNB, 3HNY, 3HOB - PubMed Abstract:
Factor VIII (FVIII) plays a critical role in blood coagulation by forming the tenase complex with factor IXa and calcium ions on a membrane surface containing negatively charged phospholipids. The tenase complex activates factor X during blood coagulation. The carboxyl-terminal C2 domain of FVIII is the main membrane-binding and von Willebrand factor-binding region of the protein. Mutations of FVIII cause hemophilia A, whereas elevation of FVIII activity is a risk factor for thromboembolic diseases. The C2 domain-membrane interaction has been proposed as a target of intervention for regulation of blood coagulation. A number of molecules that interrupt FVIII or factor V (FV) binding to cell membranes have been identified through high throughput screening or structure-based design. We report crystal structures of the FVIII C2 domain under three new crystallization conditions, and a high resolution (1.15 A) crystal structure of the FVIII C2 domain bound to a small molecular inhibitor. The latter structure shows that the inhibitor binds to the surface of an exposed beta-strand of the C2 domain, Trp(2313)-His(2315). This result indicates that the Trp(2313)-His(2315) segment is an important constituent of the membrane-binding motif and provides a model to understand the molecular mechanism of the C2 domain membrane interaction.
- State Key Laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, Chinese Academy of Sciences, Fuzhou, Fujian 350002, China.
Organizational Affiliation: