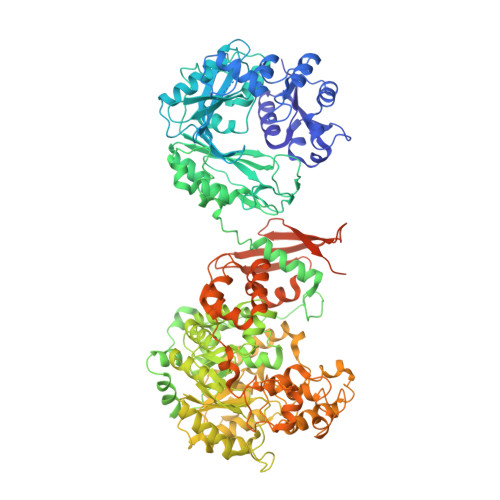
A Symmetrical Tetramer for S. aureus Pyruvate Carboxylase in Complex with Coenzyme A.
Yu, L.P., Xiang, S., Lasso, G., Gil, D., Valle, M., Tong, L.(2009) Structure 17: 823-832
- PubMed: 19523900 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.04.008
- Primary Citation Related Structures:
3HB9, 3HBL, 3HO8 - PubMed Abstract:
Pyruvate carboxylase (PC) is a conserved metabolic enzyme with important cellular functions. We report crystallographic and cryo-electron microscopy (EM) studies of Staphylococcus aureus PC (SaPC) in complex with acetyl-CoA, an allosteric activator, and mutagenesis, biochemical, and structural studies of the biotin binding site of its carboxyltransferase (CT) domain. The disease-causing A610T mutation abolishes catalytic activity by blocking biotin binding to the CT active site, and Thr908 might play a catalytic role in the CT reaction. The crystal structure of SaPC in complex with CoA reveals a symmetrical tetramer, with one CoA molecule bound to each monomer, and cryo-EM studies confirm the symmetrical nature of the tetramer. These observations are in sharp contrast to the highly asymmetrical tetramer of Rhizobium etli PC in complex with ethyl-CoA. Our structural information suggests that acetyl-CoA promotes a conformation for the dimer of the biotin carboxylase domain of PC that might be catalytically more competent.
- Department of Biological Sciences, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: