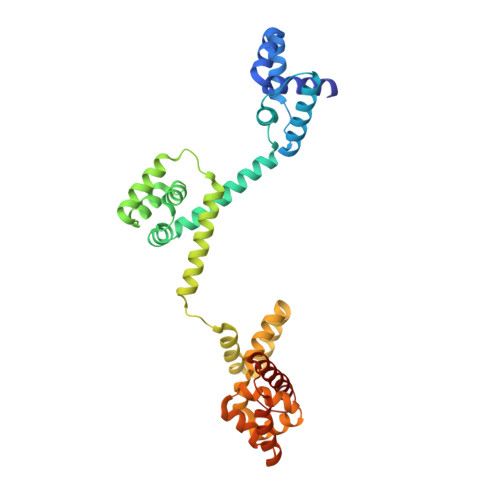
Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching
Lee, L.K., Ginsburg, M.A., Crovace, C., Donohoe, M., Stock, D.(2010) Nature 466: 996-1000
- PubMed: 20676082 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09300
- Primary Citation Related Structures:
3HJL - PubMed Abstract:
The flagellar motor drives the rotation of flagellar filaments at hundreds of revolutions per second, efficiently propelling bacteria through viscous media. The motor uses the potential energy from an electrochemical gradient of cations across the cytoplasmic membrane to generate torque. A rapid switch from anticlockwise to clockwise rotation determines whether a bacterium runs smoothly forward or tumbles to change its trajectory. A protein called FliG forms a ring in the rotor of the flagellar motor that is involved in the generation of torque through an interaction with the cation-channel-forming stator subunit MotA. FliG has been suggested to adopt distinct conformations that induce switching but these structural changes and the molecular mechanism of switching are unknown. Here we report the molecular structure of the full-length FliG protein, identify conformational changes that are involved in rotational switching and uncover the structural basis for the formation of the FliG torque ring. This allows us to propose a model of the complete ring and switching mechanism in which conformational changes in FliG reverse the electrostatic charges involved in torque generation.
- Structural and Computational Biology Division, The Victor Chang Cardiac Research Institute, Lowy Packer Building, 405 Liverpool Street, Darlinghurst, New South Wales 2010, Australia.
Organizational Affiliation: