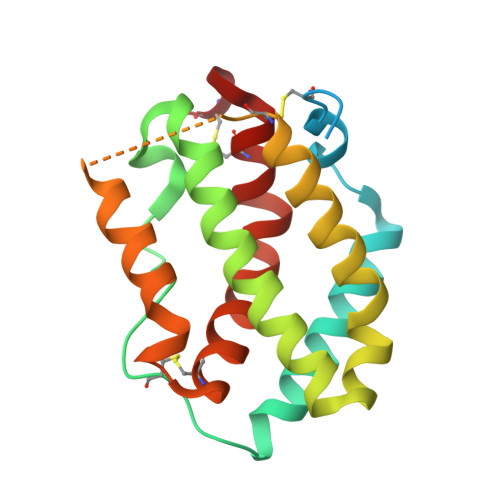
Interferon-{lambda} Is Functionally an Interferon but Structurally Related to the Interleukin-10 Family
Gad, H.H., Dellgren, C., Hamming, O.J., Vends, S., Paludan, S.R., Hartmann, R.(2009) J Biological Chem 284: 20869-20875
- PubMed: 19457860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.002923
- Primary Citation Related Structures:
3HHC - PubMed Abstract:
Interferon-lambda (IFN-lambda) is an antiviral cytokine that signals through a distinct receptor complex, composed of the IFN-lambdaR1 and interleukin-10R2 (IL-10R2) receptor chains. We have determined the crystal structure of human IFN-lambda3 and characterized the interaction with its receptor complex through structure-based site-directed mutagenesis. The ability of IFN-lambda3 mutants to signal was determined by measuring the antiviral activity and induced STAT2 phosphorylation. In conclusion, our data show that, although IFN-lambda is functionally an interferon, it is clearly structurally related to members of the IL-10 family. In particular, we found an interesting similarity between IFN-lambda and IL-22, and we suggest that IFN-lambda and IL-22 possess parallel functions, protecting epithelial tissue against viral and bacterial infections, respectively.
- Centre for Structural Biology, Department of Molecular Biology, Aarhus University, Gustav, Wieds Vej 10, 8000 Arhus C, Denmark.
Organizational Affiliation: