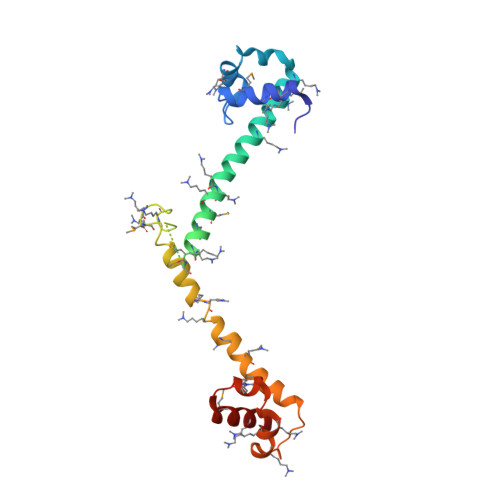
Crystal Structure of the Three Tandem FF Domains of the Transcription Elongation Regulator CA150.
Lu, M., Yang, J., Ren, Z., Sabui, S., Espejo, A., Bedford, M.T., Jacobson, R.H., Jeruzalmi, D., McMurray, J.S., Chen, X.(2009) J Mol Biology 393: 397-408
- PubMed: 19660470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.07.086
- Primary Citation Related Structures:
3HFH - PubMed Abstract:
FF domains are small protein-protein interaction modules that have two flanking conserved phenylalanine residues. They are present in proteins involved in transcription, RNA splicing, and signal transduction, and often exist in tandem arrays. Although several individual FF domain structures have been determined by NMR, the tandem nature of most FF domains has not been revealed. Here we report the 2.7-A-resolution crystal structure of the first three FF domains of the human transcription elongation factor CA150. Each FF domain is composed of three alpha-helices and a 3(10) helix between alpha-helix 2 and alpha-helix 3. The most striking feature of the structure is that an FF domain is connected to the next by an alpha-helix that continues from helix 3 to helix 1 of the next. The consequent elongated arrangement allows exposure of many charged residues within the region that can be engaged in interaction with other molecules. Binding studies using a peptide ligand suggest that a specific conformation of the FF domains might be required to achieve higher-affinity binding. Additionally, we explore potential DNA binding of the FF construct used in this study. Overall, we provide the first crystal structure of an FF domain and insights into the tandem nature of the FF domains and suggest that, in addition to protein binding, FF domains might be involved in DNA binding.
- Department of Biochemistry and Molecular Biology, University of Texas M. D. Anderson Cancer Center, Houston, 77030, USA.
Organizational Affiliation: