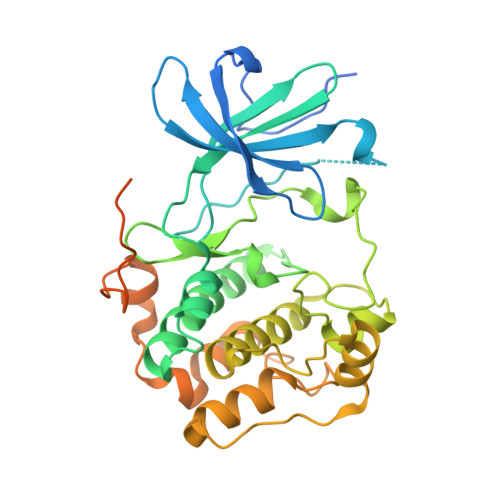
Design and synthesis of orally bioavailable serum and glucocorticoid-regulated kinase 1 (SGK1) inhibitors.
Hammond, M., Washburn, D.G., Hoang, H.T., Manns, S., Frazee, J.S., Nakamura, H., Patterson, J.R., Trizna, W., Wu, C., Azzarano, L.M., Nagilla, R., Nord, M., Trejo, R., Head, M.S., Zhao, B., Smallwood, A.M., Hightower, K., Laping, N.J., Schnackenberg, C.G., Thompson, S.K.(2009) Bioorg Med Chem Lett 19: 4441-4445
- PubMed: 19497745 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.05.051
- Primary Citation Related Structures:
3HDM, 3HDN - PubMed Abstract:
The lead serum and glucocorticoid-related kinase 1 (SGK1) inhibitors 4-(5-phenyl-1H-pyrrolo[2,3-b]pyridin-3-yl)benzoic acid (1) and {4-[5-(2-naphthalenyl)-1H-pyrrolo[2,3-b]pyridin-3-yl]phenyl}acetic acid (2) suffer from low DNAUC values in rat, due in part to formation and excretion of glucuronic acid conjugates. These PK/glucuronidation issues were addressed either by incorporating a substituent on the 3-phenyl ring ortho to the key carboxylate functionality of 1 or by substituting on the group in between the carboxylate and phenyl ring of 2. Three of these analogs have been identified as having good SGK1 inhibition potency and have DNAUC values suitable for in vivo testing.
- Department of Chemistry, Metabolic Pathways Centre for Excellence in Drug Discovery, GlaxoSmithKline Pharmaceuticals, 709 Swedeland Road, King of Prussia, PA 19406, USA. marlys.2.hammond@gsk.com
Organizational Affiliation: