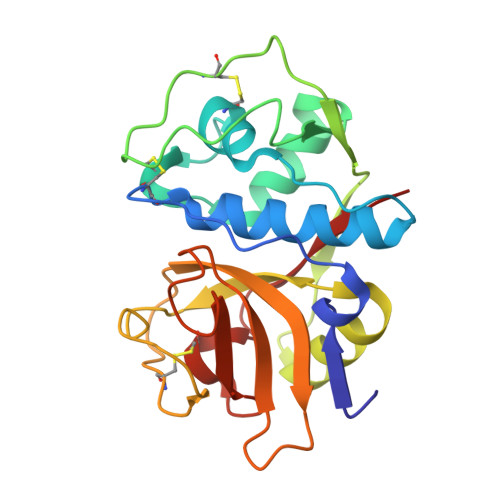
Novel non-peptidic vinylsulfones targeting the S2 and S3 subsites of parasite cysteine proteases.
Bryant, C., Kerr, I.D., Debnath, M., Ang, K.K., Ratnam, J., Ferreira, R.S., Jaishankar, P., Zhao, D., Arkin, M.R., McKerrow, J.H., Brinen, L.S., Renslo, A.R.(2009) Bioorg Med Chem Lett 19: 6218-6221
- PubMed: 19773167 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2009.08.098
- Primary Citation Related Structures:
3HD3 - PubMed Abstract:
We describe here the identification of non-peptidic vinylsulfones that inhibit parasite cysteine proteases in vitro and inhibit the growth of Trypanosoma brucei brucei parasites in culture. A high resolution (1.75 A) co-crystal structure of 8a bound to cruzain reveals how the non-peptidic P2/P3 moiety in such analogs bind the S2 and S3 subsites of the protease, effectively recapitulating important binding interactions present in more traditional peptide-based protease inhibitors and natural substrates.
- Small Molecule Discovery Center, University of California, San Francisco, CA 94158, USA.
Organizational Affiliation: