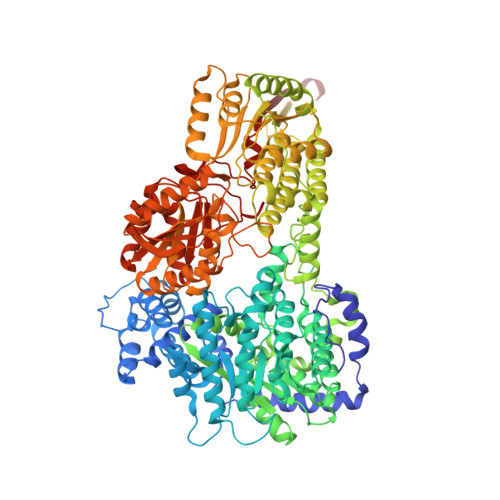
Crystal structure of the bifunctional proline utilization A flavoenzyme from Bradyrhizobium japonicum
Srivastava, D., Schuermann, J.P., White, T.A., Krishnan, N., Sanyal, N., Hura, G.L., Tan, A., Henzl, M.T., Becker, D.F., Tanner, J.J.(2010) Proc Natl Acad Sci U S A 107: 2878-2883
- PubMed: 20133651 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0906101107
- Primary Citation Related Structures:
3HAZ - PubMed Abstract:
The bifunctional proline catabolic flavoenzyme, proline utilization A (PutA), catalyzes the oxidation of proline to glutamate via the sequential activities of FAD-dependent proline dehydrogenase (PRODH) and NAD(+)-dependent Delta(1)-pyrroline-5-carboxylate dehydrogenase (P5CDH) domains. Although structures for some of the domains of PutA are known, a structure for the full-length protein has not previously been solved. Here we report the 2.1 A resolution crystal structure of PutA from Bradyrhizobium japonicum, along with data from small-angle x-ray scattering, analytical ultracentrifugation, and steady-state and rapid-reaction kinetics. PutA forms a ring-shaped tetramer in solution having a diameter of 150 A. Within each protomer, the PRODH and P5CDH active sites face each other at a distance of 41 A and are connected by a large, irregularly shaped cavity. Kinetics measurements show that glutamate production occurs without a lag phase, suggesting that the intermediate, Delta(1)-pyrroline-5-carboxylate, is preferably transferred to the P5CDH domain rather than released into the bulk medium. The structural and kinetic data imply that the cavity serves both as a microscopic vessel for the hydrolysis of Delta(1)-pyrroline-5-carboxylate to glutamate semialdehyde and a protected conduit for the transport of glutamate semialdehyde to the P5CDH active site.
- Department of Chemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: