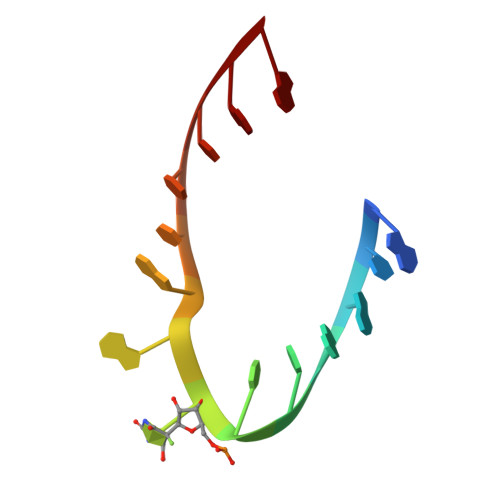
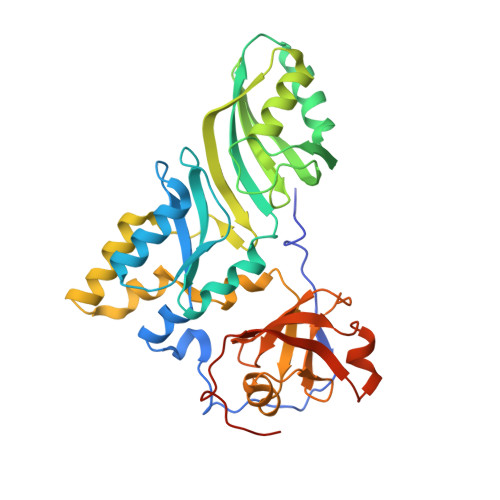
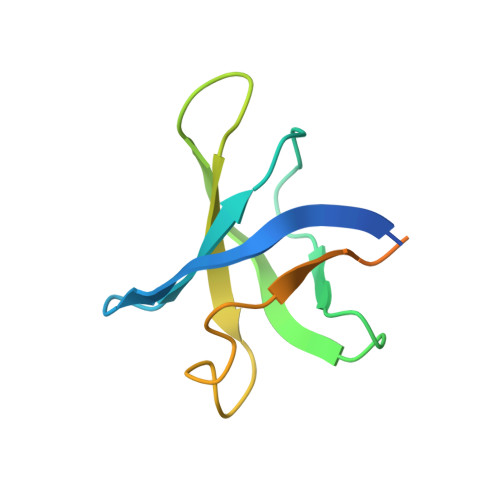
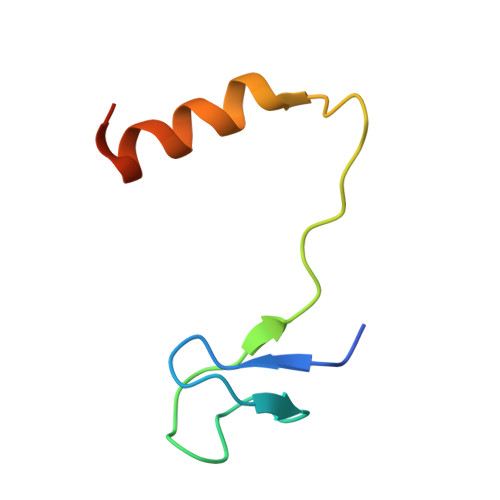
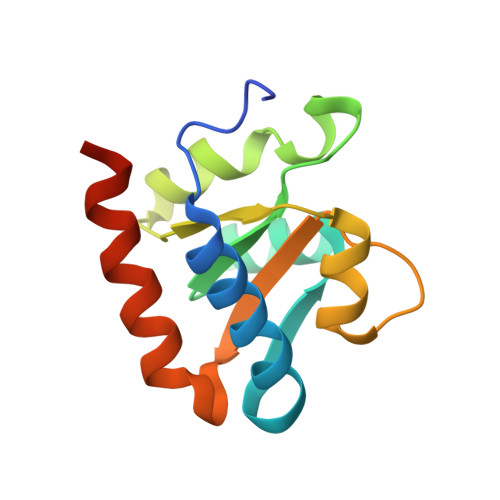
Structural mechanism of substrate RNA recruitment in H/ACA RNA-guided pseudouridine synthase.
Duan, J., Li, L., Lu, J., Wang, W., Ye, K.(2009) Mol Cell 34: 427-439
- PubMed: 19481523 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2009.05.005
- Primary Citation Related Structures:
3HAX, 3HAY - PubMed Abstract:
H/ACA RNAs form ribonucleoprotein complex (RNP) with proteins Cbf5, Nop10, L7Ae, and Gar1 and guide site-specific conversion of uridine into pseudouridine in cellular RNAs. The crystal structures of H/ACA RNP with substrate bound at the active site cleft reveal that the substrate is recruited through sequence-specific pairing with guide RNA and essential protein contacts. Substrate binding leads to a reorganization of a preset pseudouridylation pocket and an adaptive movement of the PUA domain and the lower stem of the H/ACA RNA. Moreover, a thumb loop flips from the Gar1-bound state in the substrate-free RNP structure to tightly associate with the substrate. Mutagenesis and enzyme kinetics analysis suggest a critical role of Gar1 and the thumb in substrate turnover, particularly in product release. Comparison with tRNA Psi55 synthase TruB reveals the structural conservation and adaptation between an RNA-guided and stand-alone pseudouridine synthase and provides insight into the guide-independent activity of Cbf5.
- College of Life Sciences, Peking University, Beijing, China.
Organizational Affiliation: