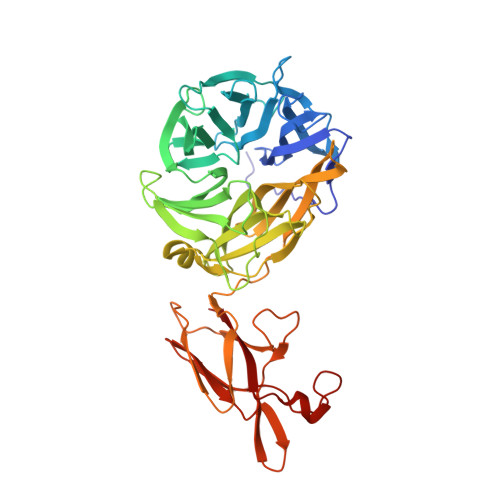
Crystal structures of respiratory pathogen neuraminidases
Hsiao, Y.-S., Parker, D., Ratner, A.J., Prince, A., Tong, L.(2009) Biochem Biophys Res Commun 380: 467-471
- PubMed: 19284989 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2009.01.108
- Primary Citation Related Structures:
3H6J, 3H71, 3H72, 3H73 - PubMed Abstract:
Currently there is pressing need to develop novel therapeutic agents for the treatment of infections by the human respiratory pathogens Pseudomonas aeruginosa and Streptococcus pneumoniae. The neuraminidases of these pathogens are important for host colonization in animal models of infection and are attractive targets for drug discovery. To aid in the development of inhibitors against these neuraminidases, we have determined the crystal structures of the P. aeruginosa enzyme NanPs and S. pneumoniae enzyme NanA at 1.6 and 1.7A resolution, respectively. In situ proteolysis with trypsin was essential for the crystallization of our recombinant NanA. The active site regions of the two enzymes are strikingly different. NanA contains a deep pocket that is similar to that in canonical neuraminidases, while the NanPs active site is much more open. The comparative studies suggest that NanPs may not be a classical neuraminidase, and may have distinct natural substrates and physiological functions. This work represents an important step in the development of drugs to prevent respiratory tract colonization by these two pathogens.
- Department of Biological Sciences, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: